
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 13,413,178 – 13,413,270 |
| Length | 92 |
| Max. P | 0.636713 |

| Location | 13,413,178 – 13,413,270 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 77.32 |
| Mean single sequence MFE | -27.47 |
| Consensus MFE | -18.97 |
| Energy contribution | -20.00 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636713 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
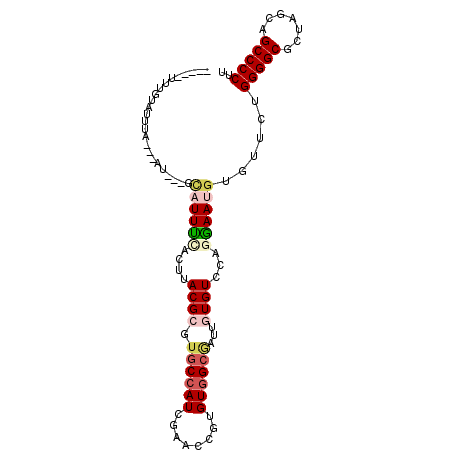
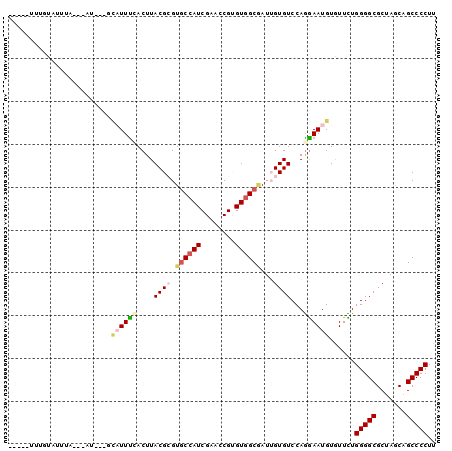
>2R_DroMel_CAF1 13413178 92 - 20766785 -----UUUGCAUUGGCAUGG---GCAUUUCACUUACGGGUGCCAUCGAACCGGGUGGCGAUGGUGUCCAGGAAUGUGUUCUGGGGCGCUAGCAGCCCCUU -----..(((....))).((---(((((((.((...(((..((((((..((....))))))))..)))))))).)))))).(((((.......))))).. ( -35.20) >DroGri_CAF1 22931 97 - 1 AAAUGUUUGUGUAUA---AUAUAUCUUUUGACUUACGCGUGCCAUCGAACCGUGUGGCAAUUGUGUCCAGGAAUGUGUUCUGGGGCGCGAGCAGCCCCUU ...(((((((((...---................((((.((((((........))))))...))))((((((.....)))))).)))))))))....... ( -27.60) >DroWil_CAF1 18734 92 - 1 -----UUUAUGUUUUUUUGU---GUUUUUCACUUACGCGUUCCAUCGAAGCGUGUGGCAAUUGUGUCCAAGAAUGUAUUUUGGGGCGCUAGCAGCCCCUU -----..(((((((((..((---(.....)))(((((((((.......)))))))))...........)))))))))....(((((.......))))).. ( -22.80) >DroMoj_CAF1 16244 89 - 1 -----UUUAUAUUUA---AU---UCAUUCUACUUACGCGUGCCAUCGAAGCGUGUAGCGAUUGUGUCCAGGAAUGUAUUUUGGGGCGCUAACAGCCCCUU -----..(((((((.---..---.(((.....((((((((.........)))))))).....))).....)))))))....(((((.......))))).. ( -23.90) >DroAna_CAF1 10411 92 - 1 -----UUAGCAUUGGCAUUA---GCAUUUCACUUACGAGUGCCAUCGAAGCGGGUGGCGAUUGUGUCCAGGAAUGUGUUCUGGGGCGCUAGCAGCCCCUU -----...((....))....---(((((((...(((((.(((((((......))))))).)))))....))))))).....(((((.......))))).. ( -32.60) >DroPer_CAF1 11160 75 - 1 -------------------------AUUUCACUUACGCGUGCCAUCGAACCGUGUGGCUAUGGUGUCCAGAAAUGUGUUUUGGGGCGCUAGCAGCCCCUU -------------------------.........(((((.(........))))))((((.((((((((.((((....)))).))))))))..)))).... ( -22.70) >consensus _____UUUGUAUUUA___AU___GCAUUUCACUUACGCGUGCCAUCGAACCGUGUGGCGAUUGUGUCCAGGAAUGUGUUCUGGGGCGCUAGCAGCCCCUU ........................((((((....((((.((((((........))))))...))))...))))))......(((((.......))))).. (-18.97 = -20.00 + 1.03)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:10:52 2006