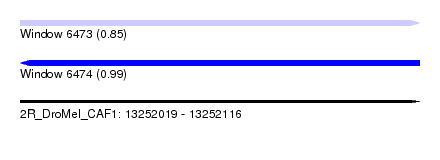

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 13,252,019 – 13,252,116 |
| Length | 97 |
| Max. P | 0.987598 |

| Location | 13,252,019 – 13,252,116 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 79.22 |
| Mean single sequence MFE | -25.71 |
| Consensus MFE | -14.55 |
| Energy contribution | -16.67 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853351 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
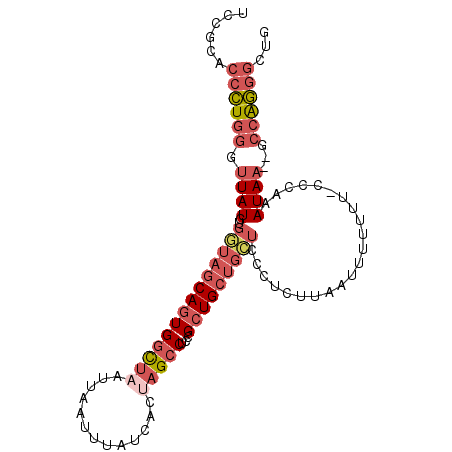
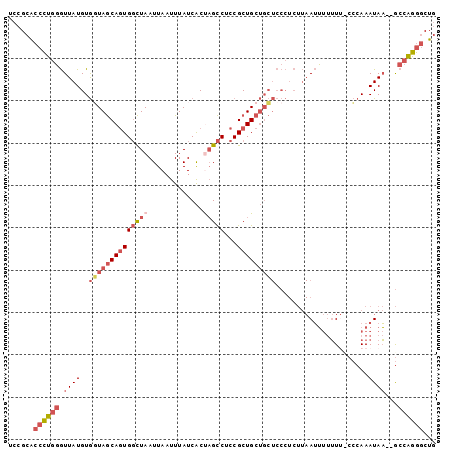
>2R_DroMel_CAF1 13252019 97 + 20766785 UCCGCACCCUGGGUUAUGUGGUAGCAGUGGCUAAUUAAUUUAUCACUAGCCUCCGCUGCUGCUCCCUCUUAAUUUUUU--CCCAAAUAA--GCCAGGGCUG ......((((((.(((((..(((((((((((((.............)))))...))))))))..).............--.....))))--.))))))... ( -30.68) >DroPse_CAF1 5415 88 + 1 GCCGCGAAUUAUAUUAUGUAUUAACAAUGCUUAAUUAAUUUAUCGCCCGUCCCCGCUGCACA---------AGACCAU--CAGAAAUAU--ACCGAAACUA ((.((((((((.((((.((((.....)))).))))))))).............))).))...---------.......--.........--.......... ( -7.61) >DroSec_CAF1 3623 99 + 1 UCCGCACCCUGGGUUAUGUGCUAGCAGUGGCUAAUUAAUUUAUCACUAGCCUCCGCUGCUGCUCCCUCUUAAUUUUUUUCCCCAAAUAA--GCCAGGGCUG ......((((((.((((..((.(((((((((((.............)))))...)))))))).......................))))--.))))))... ( -27.93) >DroSim_CAF1 3616 99 + 1 UCCGCACCCUGGGUUAUGUGGUAGCAGUGGCUAAUUAAUUUAUCACUAGCCUCCGCUGCUGCUCCCUCUUAAUUUUUUUCCCCAAAUAA--GCCAGGGCUG ......((((((.(((((..(((((((((((((.............)))))...))))))))..)....................))))--.))))))... ( -30.57) >DroYak_CAF1 3595 101 + 1 UCCGCACCCUGGGUUAUGUGGUAGCAGUGGCUAAUUAAUUUAUCACCAGCCUCCGCUGCUGCUCCCUCUUAAUUUUUUUUCCCAAAUAAGGGCCAGGGCUG ......((((((.....(..(((((.(.((((...............)))).).)))))..)..................(((......)))))))))... ( -31.76) >consensus UCCGCACCCUGGGUUAUGUGGUAGCAGUGGCUAAUUAAUUUAUCACUAGCCUCCGCUGCUGCUCCCUCUUAAUUUUUUU_CCCAAAUAA__GCCAGGGCUG ......((((((.((((..((((((((((((((.............)))))...)))))))))......................))))...))))))... (-14.55 = -16.67 + 2.12)

| Location | 13,252,019 – 13,252,116 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 79.22 |
| Mean single sequence MFE | -30.40 |
| Consensus MFE | -22.32 |
| Energy contribution | -23.28 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.73 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987598 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
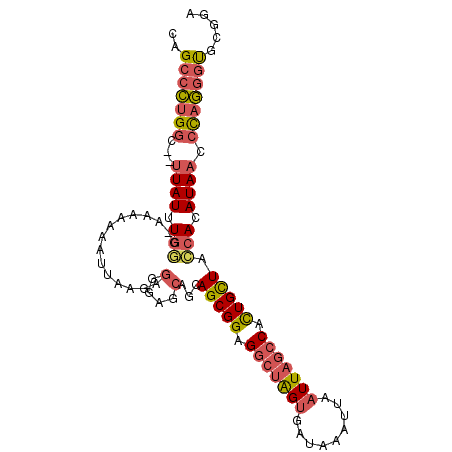
>2R_DroMel_CAF1 13252019 97 - 20766785 CAGCCCUGGC--UUAUUUGGG--AAAAAAUUAAGAGGGAGCAGCAGCGGAGGCUAGUGAUAAAUUAAUUAGCCACUGCUACCACAUAACCCAGGGUGCGGA ..(((((((.--((((.(((.--............(....)...(((((.(((((((.........))))))).))))).))).)))).)))))))..... ( -35.60) >DroPse_CAF1 5415 88 - 1 UAGUUUCGGU--AUAUUUCUG--AUGGUCU---------UGUGCAGCGGGGACGGGCGAUAAAUUAAUUAAGCAUUGUUAAUACAUAAUAUAAUUCGCGGC ..((..((((--(((((..((--(..((((---------(((((..((....)).)))..........)))).))..)))......))))))..)))..)) ( -12.50) >DroSec_CAF1 3623 99 - 1 CAGCCCUGGC--UUAUUUGGGGAAAAAAAUUAAGAGGGAGCAGCAGCGGAGGCUAGUGAUAAAUUAAUUAGCCACUGCUAGCACAUAACCCAGGGUGCGGA ..((((((((--((.(((...((......))..))).)))).(((((((.(((((((.........))))))).))))).))........))))))..... ( -33.50) >DroSim_CAF1 3616 99 - 1 CAGCCCUGGC--UUAUUUGGGGAAAAAAAUUAAGAGGGAGCAGCAGCGGAGGCUAGUGAUAAAUUAAUUAGCCACUGCUACCACAUAACCCAGGGUGCGGA ..(((((((.--((((.(((...............(....)...(((((.(((((((.........))))))).))))).))).)))).)))))))..... ( -35.60) >DroYak_CAF1 3595 101 - 1 CAGCCCUGGCCCUUAUUUGGGAAAAAAAAUUAAGAGGGAGCAGCAGCGGAGGCUGGUGAUAAAUUAAUUAGCCACUGCUACCACAUAACCCAGGGUGCGGA ..((((((((((((((((........))))...)))))......(((((.(((((((.........))))))).)))))..........)))))))..... ( -34.80) >consensus CAGCCCUGGC__UUAUUUGGG_AAAAAAAUUAAGAGGGAGCAGCAGCGGAGGCUAGUGAUAAAUUAAUUAGCCACUGCUACCACAUAACCCAGGGUGCGGA ..(((((((...((((.(((...............(....)...(((((.(((((((.........))))))).))))).))).)))).)))))))..... (-22.32 = -23.28 + 0.96)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:09:39 2006