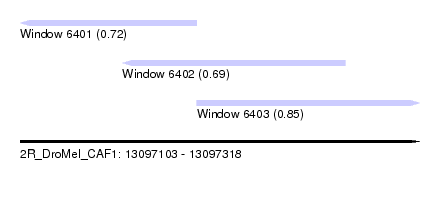
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 13,097,103 – 13,097,318 |
| Length | 215 |
| Max. P | 0.853792 |

| Location | 13,097,103 – 13,097,198 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.98 |
| Mean single sequence MFE | -30.90 |
| Consensus MFE | -27.93 |
| Energy contribution | -28.27 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 13097103 95 - 20766785 CCUUCGUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCUCGAUCGGGCCGGGUCCGACCUUUUGGAUCUUUACCGCGAUCGCGGUCAUCGCCUUC .....(((((..(((.((((((..........))))))..(((((((..((((((((....))))))))...)).))))))))...))))).... ( -32.50) >DroSim_CAF1 8304 95 - 1 CCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCGCGAUCGGGCCGGGUCCAACCUUCUGGAUCUUCACCGUGAUCGCGGUCAUCGCCUUC .....(((..(.....((((((..........))))))(((((((((..(((((((......)))))))...)).))))))))..)))....... ( -32.40) >DroYak_CAF1 8478 95 - 1 CCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAACGCGAUCGGGGCGGGUCCAACCUUCUGGAUCUUCACCGUGAUCGCGGUCCUCUCCUUC .......((.(.......((((..........)))).((((((((.((.(((((((......)))))))...)).))))))))).))........ ( -27.80) >consensus CCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCGCGAUCGGGCCGGGUCCAACCUUCUGGAUCUUCACCGUGAUCGCGGUCAUCGCCUUC .....(((..(.....((((((..........))))))(((((((((..(((((((......)))))))...)).))))))))..)))....... (-27.93 = -28.27 + 0.34)


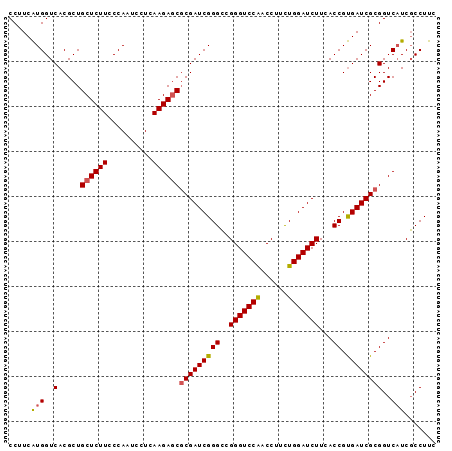
| Location | 13,097,158 – 13,097,278 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.22 |
| Mean single sequence MFE | -50.50 |
| Consensus MFE | -47.67 |
| Energy contribution | -48.67 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.692205 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 13097158 120 - 20766785 GCUGGUGAUGGCGGAACUCUUCUCGGAGGAUGUGAAGAGUGUGGCAGGAUCGAUUGCCGGCACCAGCAACUGGCUGUCGGCCUUCGUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCUC (((......))).((.(((((....((((((..((((((((((((.....(((..(((((((((((...)))).)))))))..)))..)))))...)))))))...))))))))))).)) ( -50.90) >DroSim_CAF1 8359 120 - 1 GCUGGUGAUGGCGGAGCUCUUCUCGGAGGAUGUGAAGAGUGUGGCAGGAUCGAUUGCCGGCACCAGCAACUGGCUGUCGGCCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCGC (((......)))...((((((....((((((..(((((((..(((..((((((..(((((((((((...)))).)))))))..))..))))..))))))))))...)))))))))))).. ( -53.40) >DroYak_CAF1 8533 120 - 1 GCUGGUGAUGGCGGAGCUCUUCUCGGAGGAUGUGAAGAGUGUGGCAGGAUCUAUUGCCGGCACCAGCAACUGGCUGUCGGCCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAACGC (((......)))...((..((((..((((((..(((((((..(((..((((....(((((((((((...)))).)))))))......))))..))))))))))...)))))).)))).)) ( -47.20) >consensus GCUGGUGAUGGCGGAGCUCUUCUCGGAGGAUGUGAAGAGUGUGGCAGGAUCGAUUGCCGGCACCAGCAACUGGCUGUCGGCCUUCAUGGUCACGCUGCUCUUCCCAAUCCUCAAGAGCGC (((......)))...((((((....((((((..(((((((..(((..((((((..(((((((((((...)))).)))))))..))..))))..))))))))))...)))))))))))).. (-47.67 = -48.67 + 1.00)
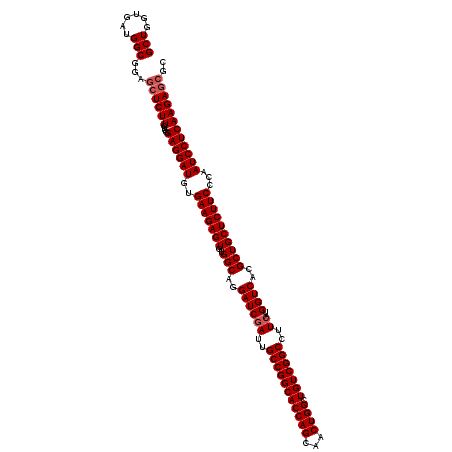


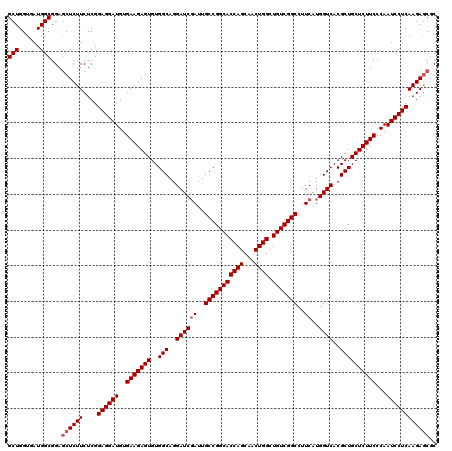
| Location | 13,097,198 – 13,097,318 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.56 |
| Mean single sequence MFE | -34.90 |
| Consensus MFE | -33.97 |
| Energy contribution | -33.97 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 13097198 120 + 20766785 CCGACAGCCAGUUGCUGGUGCCGGCAAUCGAUCCUGCCACACUCUUCACAUCCUCCGAGAAGAGUUCCGCCAUCACCAGCCACGGCACCGGUCCGAAUCCAAUCGAGAAGAAGAUUAUGA (((...(((.((.(((((((..((((........))))..(((((((.(.......).)))))))........))))))).)))))..)))(((((......))).))............ ( -36.50) >DroSim_CAF1 8399 120 + 1 CCGACAGCCAGUUGCUGGUGCCGGCAAUCGAUCCUGCCACACUCUUCACAUCCUCCGAGAAGAGCUCCGCCAUCACCAGCCACGGCACCGGUCCAAAUCCAAUCGAGAAGAAUAUAAUGA .(((..(((.((.(((((((..((((........))))...((((((.(.......).)))))).........))))))).)))))...((.......))..)))............... ( -33.90) >DroYak_CAF1 8573 120 + 1 CCGACAGCCAGUUGCUGGUGCCGGCAAUAGAUCCUGCCACACUCUUCACAUCCUCCGAGAAGAGCUCCGCCAUCACCAGCCACGGCACCGGUCCGAAUCCAAUCGAAAAGAAGACAAUAA (((...(((.((.(((((((..((((........))))...((((((.(.......).)))))).........))))))).)))))..)))..(((......)))............... ( -34.30) >consensus CCGACAGCCAGUUGCUGGUGCCGGCAAUCGAUCCUGCCACACUCUUCACAUCCUCCGAGAAGAGCUCCGCCAUCACCAGCCACGGCACCGGUCCGAAUCCAAUCGAGAAGAAGAUAAUGA .(((..(((.((.(((((((..((((........))))...((((((.(.......).)))))).........))))))).)))))...((.......))..)))............... (-33.97 = -33.97 + -0.00)

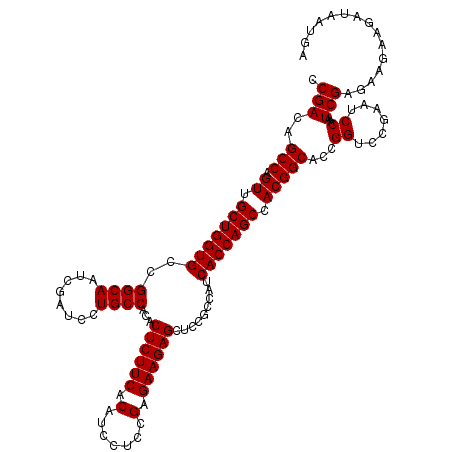
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:08:30 2006