
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,958,166 – 12,958,268 |
| Length | 102 |
| Max. P | 0.650754 |

| Location | 12,958,166 – 12,958,268 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 73.28 |
| Mean single sequence MFE | -30.11 |
| Consensus MFE | -17.34 |
| Energy contribution | -17.90 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.650754 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
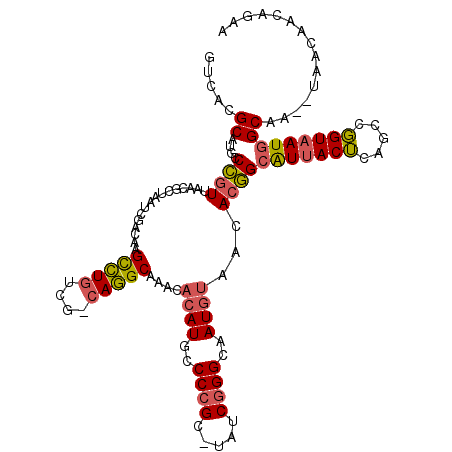
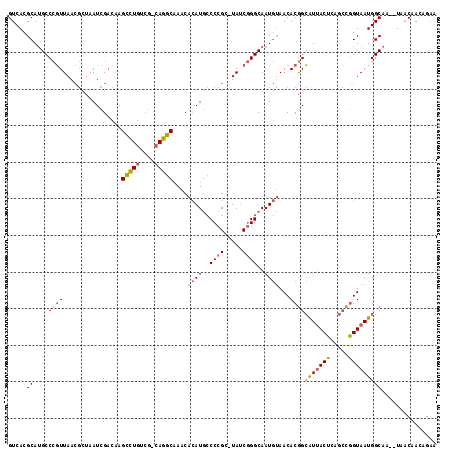
>2R_DroMel_CAF1 12958166 102 - 20766785 GUCACGCACGCCUUUUAA-----AUAGUCCAGUGUGU---AAUGCCAUCUUAUCGCCCGCUUAACCGGCAAU--AACACAGAAUAACA---AUUGUAAUGGCAACUUGGCAACAG-- (((((((((.........-----........))))))---..((((((.((((.(((.(.....).))).))--)).((((.......---.)))).))))))....))).....-- ( -20.43) >DroPse_CAF1 9359 114 - 1 GUCACGCAUGCCCGUUAACGCUAAUCGACAAGCCUGUCGUCAGGCAAACACAUGCCCCGC-UAUCGGGCAAUGUAACACGGCGUUACUCAGCCGGUAAUGGCAA--UAACAACAGAA ((((.(((((((((.((.((.....(((((....)))))...((((......)))).)).-)).))))).)))).((.((((........))))))..))))..--........... ( -35.40) >DroPer_CAF1 9392 114 - 1 GUCACGCAUGCCCGUUAACGCUAAUCGACAAGCCUGUCGCCAGGCAAACACAUGCCCCGC-UAUCGGGCAAUGUAACACGGCAUUACUCAGCCGGUAAUGGCAA--UAACAACAGAA ((((.(((((((((.((.((.....(((((....)))))...((((......)))).)).-)).))))).)))).((.((((........))))))..))))..--........... ( -34.50) >consensus GUCACGCAUGCCCGUUAACGCUAAUCGACAAGCCUGUCG_CAGGCAAACACAUGCCCCGC_UAUCGGGCAAUGUAACACGGCAUUACUCAGCCGGUAAUGGCAA__UAACAACAGAA .....((....((((................(((((....)))))....((((..((((.....))))..))))...))))(((((((.....)))))))))............... (-17.34 = -17.90 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:07:37 2006