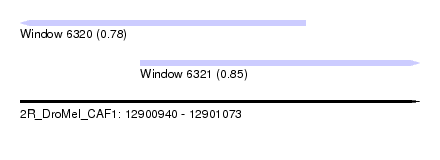
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,900,940 – 12,901,073 |
| Length | 133 |
| Max. P | 0.849161 |

| Location | 12,900,940 – 12,901,035 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 75.57 |
| Mean single sequence MFE | -35.63 |
| Consensus MFE | -19.84 |
| Energy contribution | -20.77 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.783681 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12900940 95 - 20766785 ---------------------GGGGGGAGUCUACCCCAAGUGGGUGGCUGGUGGCUUAAUGCCCAAAUUCAUGACAUACUUCAAGGCGCUGCCAGUGACCCAAGACCAAAAUUUAU ---------------------((((((......))))...(((((.(((((..((.....(((.....................)))))..))))).)))))...))......... ( -35.50) >DroSec_CAF1 24250 96 - 1 --------------------AGGUGGGGGUCUGCCCCAAGUGGGUGGCUGGUGGCUUAAUGCCCAAAUUCAUGGCAUACUUCAAGGCGUUGCCGGCGACCCAAGACCAAAAUUUAU --------------------.(((.((((....))))...(((((.(((((..((...(((((.........))))).((....)).))..))))).)))))..)))......... ( -40.30) >DroSim_CAF1 11818 97 - 1 -----------------UGGAGGAGGGGG--CGCCCCAAGUGGGUGUGUGGUGGCUUAAUGCCCAAAUUCAUGGCAUACUUCAAGGCGCUGCCGGCGACCCAAGACCAAAAUUUAU -----------------(((....((.((--((((((....))((((((.(((((.....))).......)).)))))).....)))))).))((....))....)))........ ( -31.61) >DroEre_CAF1 26301 85 - 1 -------------------------------UGCCCCAAGUGGGUGGUUGGUGGCUUAAUGCCCAAAUUCAUGGCAUACUUCAAGGCGCUGCCGGCGACCCAAGACCGAAAUUUAU -------------------------------.........(((((.(((((..((...(((((.........))))).((....)).))..))))).))))).............. ( -27.60) >DroYak_CAF1 25452 106 - 1 CUCGUCAGU--------UGGGUGGCAUGG--CCCCCCAAGUGGAUGGUUGCUGGCUUAAUGCCGAAAUUCAUGGCAUACUUCAAGGCGCUGCCGGCGACCCAAGACCGAAAUUUAU .((((((.(--------((((.(((...)--)).))))).)))..((((((((((...((((((.......)))))).((....))....))))))))))......)))....... ( -39.00) >DroAna_CAF1 24695 110 - 1 GUCCCCACCGGGCAGGGCUUUGGAGUGGG--UGGGUCUAGUG----GGCGGUGGCUUAAUGUCCAAAUUCAUGGCAUACUUCAAGGCGCUGCCGACGACCCUAGACCGAAAUUUAU (((((((((((((...))))..).)))))--.(((((..((.----(((((((.(((.(((.(((......)))))).....))).))))))).)))))))..))).......... ( -39.80) >consensus _____________________GGAGGGGG__UGCCCCAAGUGGGUGGCUGGUGGCUUAAUGCCCAAAUUCAUGGCAUACUUCAAGGCGCUGCCGGCGACCCAAGACCAAAAUUUAU ........................................(((((.(((((((((...(((((.........))))).((....)).))))))))).))))).............. (-19.84 = -20.77 + 0.92)
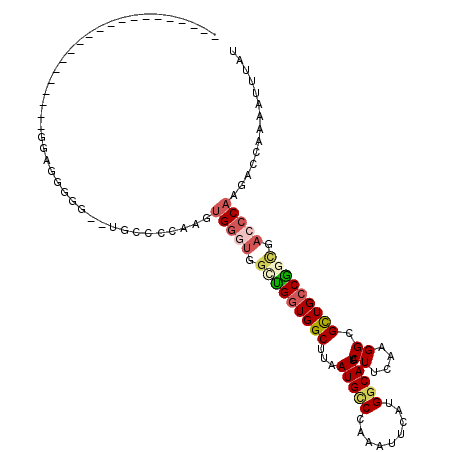


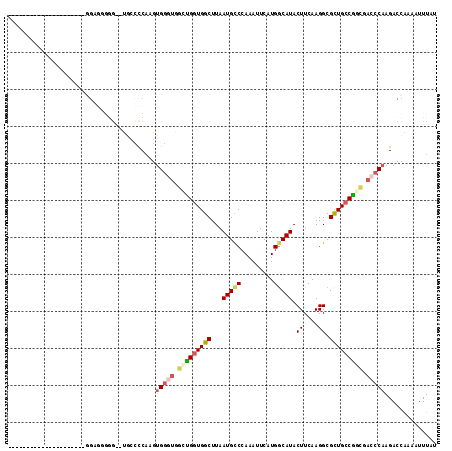
| Location | 12,900,980 – 12,901,073 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 78.98 |
| Mean single sequence MFE | -23.18 |
| Consensus MFE | -15.40 |
| Energy contribution | -14.96 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12900980 93 + 20766785 UGUCAUGAAUUUGGGCAUUAAGCCACCAGCCACCCACUUGGGGUAGACUCCCCCC---------------CCCCUUCAUUUUCCUAGACAAUUUGAGGCAAUCAUGCA .(.(((((.....(((.....)))....(((........((((..(......)..---------------)))).(((..((.......))..))))))..)))))). ( -21.90) >DroSec_CAF1 24290 94 + 1 UGCCAUGAAUUUGGGCAUUAAGCCACCAGCCACCCACUUGGGGCAGACCCCCACCU--------------CCCUCUCAUUUUCCUAGACAAUUUGAGGCAAUCAUGCA .((.((((.....(((.....)))....(((.......(((((.....)))))...--------------.....(((..((.......))..))))))..)))))). ( -21.90) >DroSim_CAF1 11858 97 + 1 UGCCAUGAAUUUGGGCAUUAAGCCACCACACACCCACUUGGGGCG--CCCCCUCCUCCA---------CCCCCUCUCAUUUUCCUAGACAAUUUGAGGCAAUCAUGCA .((.((((....((((.....(((.(((..........)))))))--))).........---------...((((..(((.((...)).)))..))))...)))))). ( -21.40) >DroEre_CAF1 26341 81 + 1 UGCCAUGAAUUUGGGCAUUAAGCCACCAACCACCCACUUGGGGCA---------------------------CCCCCACUUUCGUAGACAAUUUGAGGCAAUCAUGCA (((((((((..((((......(((.((((........))))))).---------------------------..))))..)))))...(.....).))))........ ( -21.30) >DroYak_CAF1 25492 106 + 1 UGCCAUGAAUUUCGGCAUUAAGCCAGCAACCAUCCACUUGGGGGG--CCAUGCCACCCAACUGACGAGCCCCCCUUCAUUUUCCUAGACAAUUUGAGGCAAUCAUGCA ((((.(((((((((((.....)))...............((((((--(..((.((......)).)).)))))))............)).)))))).))))........ ( -29.40) >consensus UGCCAUGAAUUUGGGCAUUAAGCCACCAACCACCCACUUGGGGCA__CCCCCCCC_______________CCCCCUCAUUUUCCUAGACAAUUUGAGGCAAUCAUGCA .(.(((((.....(((.....)))........(((....)))...............................(((((..((.......))..)))))...)))))). (-15.40 = -14.96 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:07:12 2006