
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,862,662 – 12,862,763 |
| Length | 101 |
| Max. P | 0.989797 |

| Location | 12,862,662 – 12,862,763 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.64 |
| Mean single sequence MFE | -44.65 |
| Consensus MFE | -28.76 |
| Energy contribution | -29.20 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.18 |
| SVM RNA-class probability | 0.989797 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
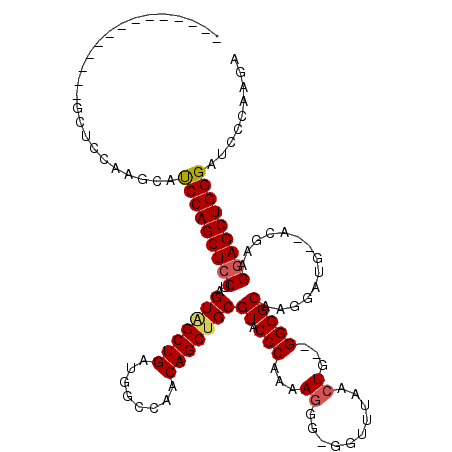
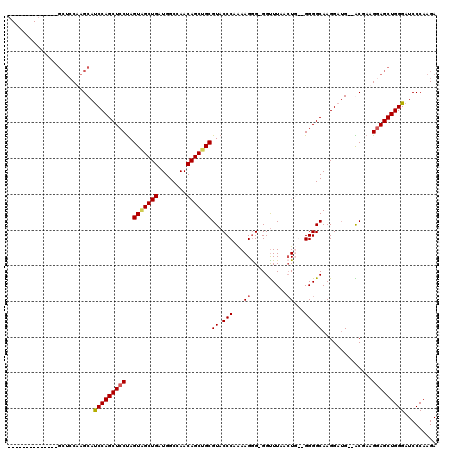
>2R_DroMel_CAF1 12862662 101 + 20766785 --------------GCUCCAAGCAUCCAGCUCCUAGUAGCUGAUGGCCAACAGCUGCGUACCCAAAAGGG-GGUUUAACUU--GGGGCAAGGAUG--ACGAUGGAGCUGGGAUCCCAAGA --------------((((((.((((((.(((((..(((((((........)))))))..((((......)-))).......--)))))..)))))--.)..))))))(((....)))... ( -44.00) >DroSim_CAF1 217971 101 + 1 --------------GCUCCAAGCAUCCAGCUCCUAGUCGCUGAUGGCCAACAGCUGCGUACCCAAAAGGG-GGUUUAAGUU--GGGGCAAGGAUG--ACGAUGGAGCUGGGAUCCCAAGA --------------((.....)).(((((((((..((((((....(((..(((((....((((......)-)))...))))--).)))..)).))--))...)))))))))......... ( -41.60) >DroEre_CAF1 210156 110 + 1 GCUUGCAGCUC-------CCAGCACCCAGCUCCCAGUAGCUGAUGGCCAACAGCUGCGUACCCAGAAGAG-GGUUUAACUGGCGGGGCAAGGAUG--GCCAAGGAGCUGGGCUCCCAAGA .((((..((..-------...)).((((((((((((...))).((((((.(.(((.(((((((......)-))).......))).)))...).))--)))).)))))))))....)))). ( -45.91) >DroYak_CAF1 216022 119 + 1 UCUUCCAGCUCCAAGCUCCAAGCAUCCAGCUCCUAGUGGCUGAUGGCCAACAGCUGCGUACCCAAAAGGGGGGCUUAACUGG-GGGGCAAGGAAGGAGCGAAGUAGCUGGGAUCCCAAGA ..(((((((((...(((((.....(((.((((((.(..((((........))))..)(..(((.....)))..).......)-)))))..))).)))))...).))))))))........ ( -47.10) >consensus ______________GCUCCAAGCAUCCAGCUCCUAGUAGCUGAUGGCCAACAGCUGCGUACCCAAAAGGG_GGUUUAACUG__GGGGCAAGGAUG__ACGAAGGAGCUGGGAUCCCAAGA ........................(((((((((..(((((((........)))))))((.(((...((..........))...)))))..............)))))))))......... (-28.76 = -29.20 + 0.44)

| Location | 12,862,662 – 12,862,763 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.64 |
| Mean single sequence MFE | -40.30 |
| Consensus MFE | -25.95 |
| Energy contribution | -26.95 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720249 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
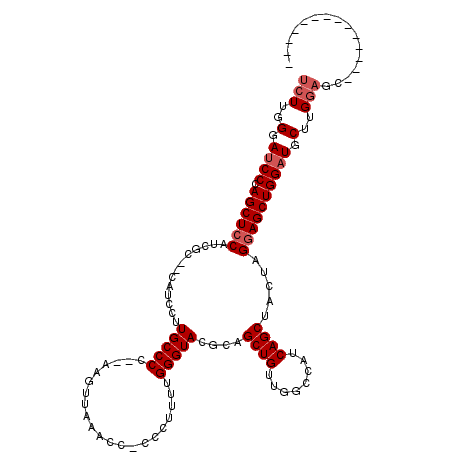
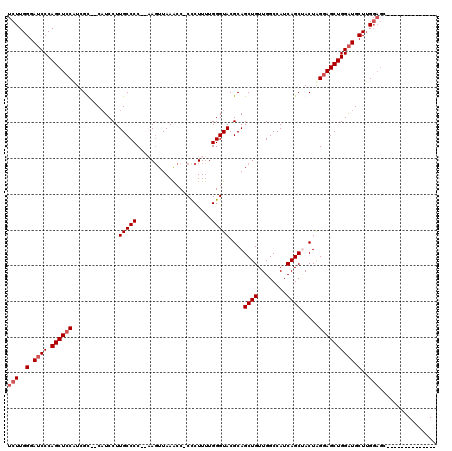
>2R_DroMel_CAF1 12862662 101 - 20766785 UCUUGGGAUCCCAGCUCCAUCGU--CAUCCUUGCCCC--AAGUUAAACC-CCCUUUUGGGUACGCAGCUGUUGGCCAUCAGCUACUAGGAGCUGGAUGCUUGGAGC-------------- ...(((....)))((((((..(.--(((((..((.((--.(((...(((-(......))))....(((((........)))))))).)).)).)))))).))))))-------------- ( -37.10) >DroSim_CAF1 217971 101 - 1 UCUUGGGAUCCCAGCUCCAUCGU--CAUCCUUGCCCC--AACUUAAACC-CCCUUUUGGGUACGCAGCUGUUGGCCAUCAGCGACUAGGAGCUGGAUGCUUGGAGC-------------- (((.((.((((.((((((...((--(.....((((((--((........-.....)))))...)))((((........)))))))..)))))))))).)).)))..-------------- ( -35.52) >DroEre_CAF1 210156 110 - 1 UCUUGGGAGCCCAGCUCCUUGGC--CAUCCUUGCCCCGCCAGUUAAACC-CUCUUCUGGGUACGCAGCUGUUGGCCAUCAGCUACUGGGAGCUGGGUGCUGG-------GAGCUGCAAGC (((..(..((((((((((.((((--((.....((...))(((((..(((-(......))))....))))).)))))).(((...))))))))))))).)..)-------))......... ( -45.60) >DroYak_CAF1 216022 119 - 1 UCUUGGGAUCCCAGCUACUUCGCUCCUUCCUUGCCCC-CCAGUUAAGCCCCCCUUUUGGGUACGCAGCUGUUGGCCAUCAGCCACUAGGAGCUGGAUGCUUGGAGCUUGGAGCUGGAAGA ..........((((((.((..(((((......((...-((((((..((..(((....)))...))..(((.((((.....)))).))).))))))..))..)))))..)))))))).... ( -43.00) >consensus UCUUGGGAUCCCAGCUCCAUCGC__CAUCCUUGCCCC__AAGUUAAACC_CCCUUUUGGGUACGCAGCUGUUGGCCAUCAGCUACUAGGAGCUGGAUGCUUGGAGC______________ (((..(.((((.((((((.............(((((.....................)))))....((((........)))).....)))))))))).)..)))................ (-25.95 = -26.95 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:31 2006