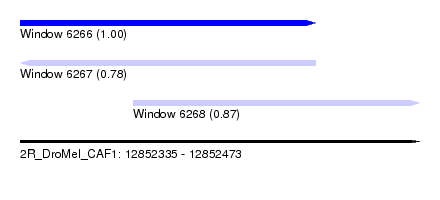

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,852,335 – 12,852,473 |
| Length | 138 |
| Max. P | 0.999902 |

| Location | 12,852,335 – 12,852,437 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 94.73 |
| Mean single sequence MFE | -29.20 |
| Consensus MFE | -26.74 |
| Energy contribution | -26.94 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.92 |
| SVM decision value | 4.46 |
| SVM RNA-class probability | 0.999902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12852335 102 + 20766785 UAGACCAGAUAUUGAUUGAACUAU-CAUUUGUGGCUGGACAUGUCUCCGGCCACAAAACAAGGCACAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ....((.(((..((((......))-))((((((((((((......))))))))))))....((.((......(((((....)).)))......))))))))). ( -29.40) >DroSec_CAF1 196091 102 + 1 UAGACCAGAUAUUGAUUGAACUAU-CAUUUGUGGCUGGACAUAUCUCCGGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ............((((......))-))((((((((((((......)))))))))))).(((.(((........)))....)))........(((.....))). ( -28.90) >DroSim_CAF1 204537 102 + 1 UAGACCAGAUAUUGAUUGAACUAU-CAUUUGUGGCCGGACAUAUUUCCUGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ............((((......))-))((((((((.(((......))).)))))))).(((.(((........)))....)))........(((.....))). ( -25.00) >DroEre_CAF1 199736 102 + 1 CAGACCAGAUAUUGAUUGAACUAU-CAUUUGUGGCUGGACAUAUCCCCGGCCACAAAACAAGGCCCAAAACAUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ....((......((((......))-))(((((((((((........)))))))))))....))........((((((....)).))))...(((.....))). ( -28.60) >DroYak_CAF1 204203 103 + 1 UAGACCAGAUAUUGAUUGAACUAUCCAUUUGUGGCUGGACAUAUCUCCGGCCACAAAACGUGGUCCAGAACAUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ..(((((((((.(......).))))..((((((((((((......))))))))))))...)))))......((((((....)).))))...(((.....))). ( -34.10) >consensus UAGACCAGAUAUUGAUUGAACUAU_CAUUUGUGGCUGGACAUAUCUCCGGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGG ...........(..((((.........((((((((((((......)))))))))))).((((........))))...))))..).......(((.....))). (-26.74 = -26.94 + 0.20)

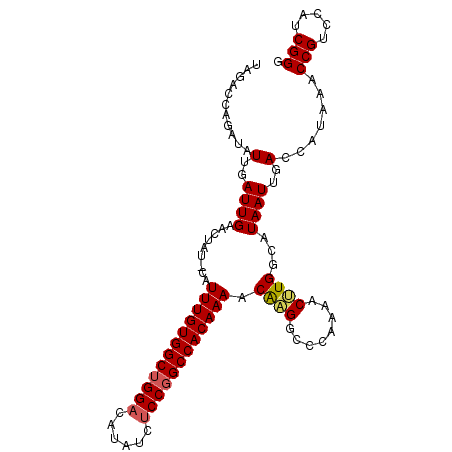
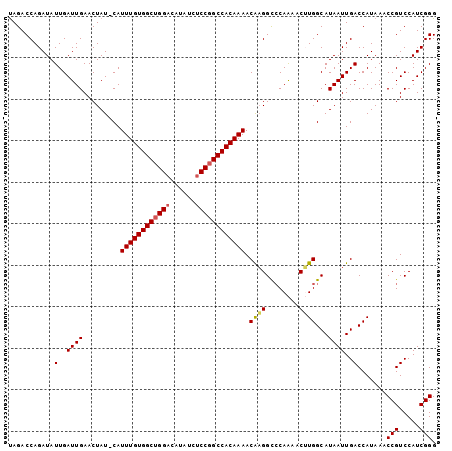
| Location | 12,852,335 – 12,852,437 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 94.73 |
| Mean single sequence MFE | -28.70 |
| Consensus MFE | -27.68 |
| Energy contribution | -27.56 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778225 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12852335 102 - 20766785 CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAAGUUUUGUGCCUUGUUUUGUGGCCGGAGACAUGUCCAGCCACAAAUG-AUAGUUCAAUCAAUAUCUGGUCUA ((.(((((((((..(.((((.......)))).)..)))).))....((((((((.(((......))).))))))))((-((......))))..))).)).... ( -25.90) >DroSec_CAF1 196091 102 - 1 CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAAGUUUUGGGCCUUGUUUUGUGGCCGGAGAUAUGUCCAGCCACAAAUG-AUAGUUCAAUCAAUAUCUGGUCUA .(((.....)))....((((.......)))).....((((((....((((((((.(((......))).))))))))((-((......))))......)))))) ( -28.00) >DroSim_CAF1 204537 102 - 1 CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAAGUUUUGGGCCUUGUUUUGUGGCAGGAAAUAUGUCCGGCCACAAAUG-AUAGUUCAAUCAAUAUCUGGUCUA .(((.....)))....((((.......)))).....((((((....((((((((.(((......))).))))))))((-((......))))......)))))) ( -28.10) >DroEre_CAF1 199736 102 - 1 CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAUGUUUUGGGCCUUGUUUUGUGGCCGGGGAUAUGUCCAGCCACAAAUG-AUAGUUCAAUCAAUAUCUGGUCUG .(((.....)))..((((((.......))))))....(((((....((((((((.(((......))).))))))))((-((......))))......))))). ( -28.70) >DroYak_CAF1 204203 103 - 1 CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAUGUUCUGGACCACGUUUUGUGGCCGGAGAUAUGUCCAGCCACAAAUGGAUAGUUCAAUCAAUAUCUGGUCUA .........(((..((((((.......))))))..)))((((....((((((((.(((......))).)))))))).(((((..........))))))))).. ( -32.80) >consensus CCCGAUGGACGGUUUAUGGUCAAUUAUGCCAAGUUUUGGGCCUUGUUUUGUGGCCGGAGAUAUGUCCAGCCACAAAUG_AUAGUUCAAUCAAUAUCUGGUCUA ...(((((((((((((((((.......)))).....))))))....((((((((.(((......))).))))))))......)))).)))............. (-27.68 = -27.56 + -0.12)

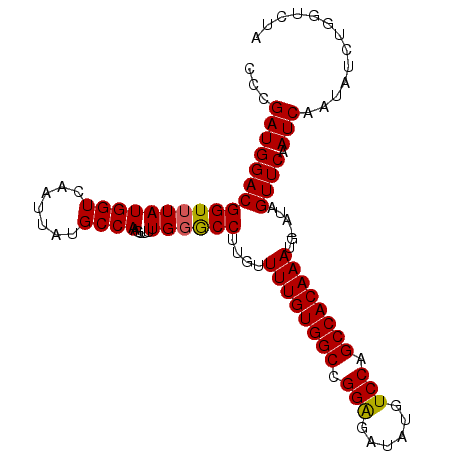
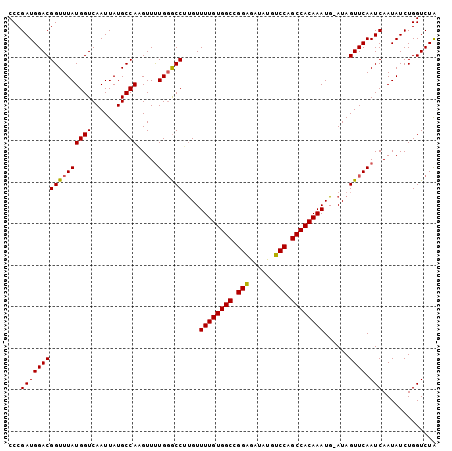
| Location | 12,852,374 – 12,852,473 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 94.95 |
| Mean single sequence MFE | -19.58 |
| Consensus MFE | -15.52 |
| Energy contribution | -16.40 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
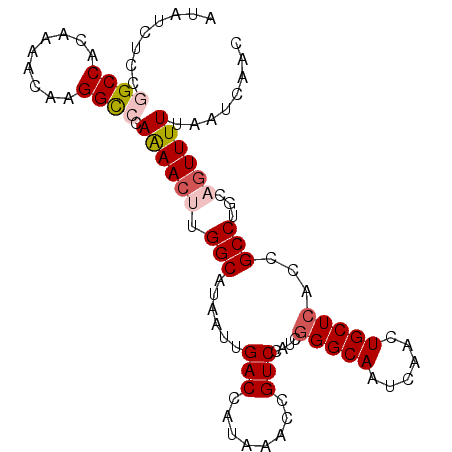
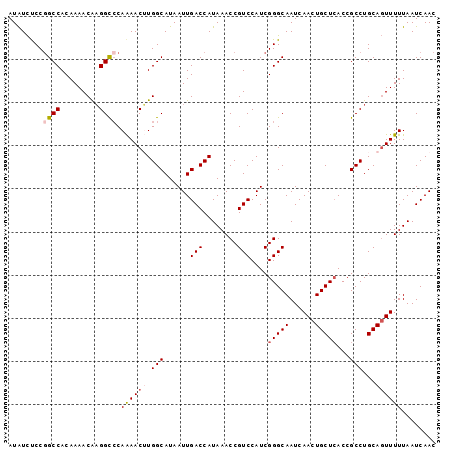
>2R_DroMel_CAF1 12852374 99 + 20766785 AUGUCUCCGGCCACAAAACAAGGCACAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUAACCGCCUGCAGUUUUUAAUCAAC .(((..((((((.........)))........(((((....)).)))............))))))...((((((.........)))))).......... ( -17.00) >DroSec_CAF1 196130 99 + 1 AUAUCUCCGGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUCACCGCCUGCAGUUUUUAAUCAAC ........((((.........)))).((((((.(((......(((........)))....(((((......)))))...)))...))))))........ ( -21.40) >DroSim_CAF1 204576 99 + 1 AUAUUUCCUGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUCACCGCCUGCACUUUUUAAUCAAC ....................((((........(((((....)).))).............(((((......)))))...))))................ ( -14.90) >DroEre_CAF1 199775 99 + 1 AUAUCCCCGGCCACAAAACAAGGCCCAAAACAUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUCACCGCCUGCAGUUUUUAAUCAAC .....(((((((.........))))......((((((....)).))))............))).....((((((.........)))))).......... ( -21.30) >DroYak_CAF1 204243 99 + 1 AUAUCUCCGGCCACAAAACGUGGUCCAGAACAUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUCACCGCCUGCAGUUUUUAAUCAAC ...(((..((((((.....)))))).)))..((((((....)).)))).....((((...))))....((((((.........)))))).......... ( -23.30) >consensus AUAUCUCCGGCCACAAAACAAGGCCCAAAACUUGGCAUAAUUGACCAUAAACCGUCCAUCGGGCAAUCAACUGCUCACCGCCUGCAGUUUUUAAUCAAC ........((((.........)))).((((((.(((......(((........)))....(((((......)))))...)))...))))))........ (-15.52 = -16.40 + 0.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:20 2006