
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,362,775 – 2,362,895 |
| Length | 120 |
| Max. P | 0.764484 |

| Location | 2,362,775 – 2,362,895 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.16 |
| Mean single sequence MFE | -38.16 |
| Consensus MFE | -29.10 |
| Energy contribution | -29.54 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.667522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
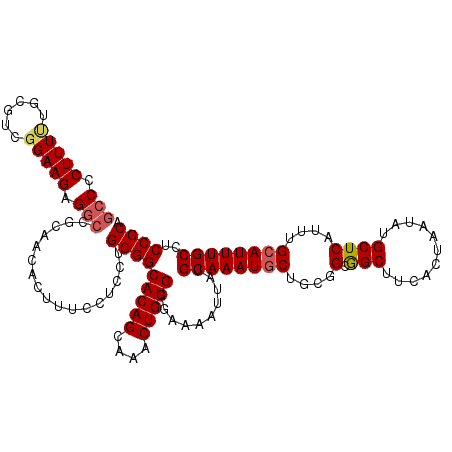
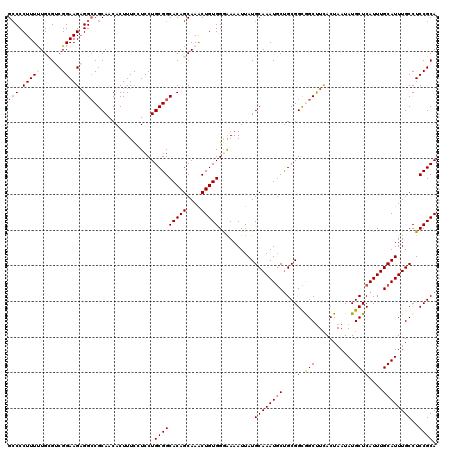
>2R_DroMel_CAF1 2362775 120 + 20766785 GACCCUUUUUGCGUCGGAAGAGGACGCAACACUUUCGUCCUGCGGCACAGCAAACUGUGGGAAAAUUAUGCAAAUGCUGCGGCGGCUUUACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA (((.....(((((((.......))))))).......)))..(((((((((....)))))((......(((((((((..(((..((.....))....))).)))))))))...)).)))). ( -35.30) >DroSec_CAF1 12288 120 + 1 GCCCCUUUUUGCGUCGGAAGAGGCCGCAACACUUUCCUCCUGCGGCACAGCAAGCUGUGGGAAAAUUAUGCAAAUGCUGCGGCGGCUUCACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA .........((((..((..((((((((..........(((..((((.......))))..)))......((((.....))))))))))))......((((......))))...))..)))) ( -42.40) >DroSim_CAF1 12399 120 + 1 GCCCCUUUUUGCGUCGGAAGAGGCCGCAACACUUUCCUCCUGCGGCACAGCAAACUGUGGGAAAAUUAUGCAAAUGCUGCGGCGGCUUCACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA .((((..(((((.((....)).((((((............))))))...)))))..).)))........((....)).((((.(((.........((((......))))..))).)))). ( -40.30) >DroEre_CAF1 12100 119 + 1 GCCCCUUUCUGAG-CGGAAGCGGCCGCAACACUUUCAUCCUGCGGCACAGCAAACUGUGGAAAAAUUAUGCAAAUGCCGCGGCAGCCUCACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA ............(-((((.(((((((((............)))).(((((....)))))................)))))(((((..........((((......)))))))))))))). ( -36.80) >DroYak_CAF1 11783 118 + 1 GCCCCUUU-UGAG-CGGAAGAGGCCGCAACACUUUUCUCCUGCGGCACAGCAAACUGUGGAGAAAUUAUGCAAAUUCUGCGGUAGCUUCACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA ........-...(-((((....((((((............))))))...(((((..((((((......((((.....))))....)))))).....(((......)))))))).))))). ( -36.00) >consensus GCCCCUUUUUGCGUCGGAAGAGGCCGCAACACUUUCCUCCUGCGGCACAGCAAACUGUGGGAAAAUUAUGCAAAUGCUGCGGCGGCUUCACUAAUAUGCUCAUUUGCAUUUGCCUCCGCA (((.(((((......))))).))).................(((((((((....)))))..........((((((((....(.(((...........))))....))))))))..)))). (-29.10 = -29.54 + 0.44)

| Location | 2,362,775 – 2,362,895 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.16 |
| Mean single sequence MFE | -40.86 |
| Consensus MFE | -32.98 |
| Energy contribution | -32.66 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.764484 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
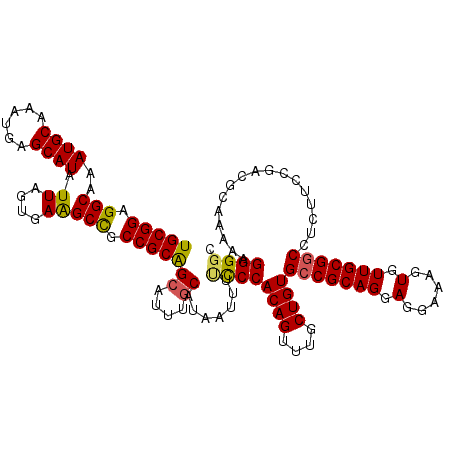
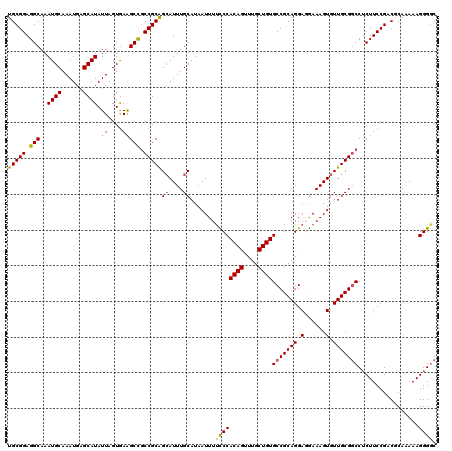
>2R_DroMel_CAF1 2362775 120 - 20766785 UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUAAAGCCGCCGCAGCAUUUGCAUAAUUUUCCCACAGUUUGCUGUGCCGCAGGACGAAAGUGUUGCGUCCUCUUCCGACGCAAAAAGGGUC (((((.(((..((((......)))).((....))))).)))))((....))........(((((((....))))((.((((.((....)).))))(((.......))))).....))).. ( -38.30) >DroSec_CAF1 12288 120 - 1 UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUGAAGCCGCCGCAGCAUUUGCAUAAUUUUCCCACAGCUUGCUGUGCCGCAGGAGGAAAGUGUUGCGGCCUCUUCCGACGCAAAAAGGGGC ((((..(....((((......))))......((((..((((((((((((......(((((((((((....))))).....))))))))))))))))))..)))))..))))......... ( -41.60) >DroSim_CAF1 12399 120 - 1 UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUGAAGCCGCCGCAGCAUUUGCAUAAUUUUCCCACAGUUUGCUGUGCCGCAGGAGGAAAGUGUUGCGGCCUCUUCCGACGCAAAAAGGGGC ((((..(....((((......))))......((((..((((((((((((......(((((((((((....))))).....))))))))))))))))))..)))))..))))......... ( -41.40) >DroEre_CAF1 12100 119 - 1 UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUGAGGCUGCCGCGGCAUUUGCAUAAUUUUUCCACAGUUUGCUGUGCCGCAGGAUGAAAGUGUUGCGGCCGCUUCCG-CUCAGAAAGGGGC .((((((((..(((((((((.((....(((.....)))..))..))))))))).........((((....))))(((((((.((....)).))))))).)))))))-)............ ( -47.50) >DroYak_CAF1 11783 118 - 1 UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUGAAGCUACCGCAGAAUUUGCAUAAUUUCUCCACAGUUUGCUGUGCCGCAGGAGAAAAGUGUUGCGGCCUCUUCCG-CUCA-AAAGGGGC .(((((((...((((((((..((....(((.....)))..))...)))))))).........((((....))))(((((((.(......).)))))))..))))))-)...-........ ( -35.50) >consensus UGCGGAGGCAAAUGCAAAUGAGCAUAUUAGUGAAGCCGCCGCAGCAUUUGCAUAAUUUUCCCACAGUUUGCUGUGCCGCAGGAGGAAAGUGUUGCGGCCUCUUCCGACGCAAAAAGGGGC (((((.(((..((((......)))).((....))))).)))))((....)).......((((((((....))))(((((((.(......).))))))).................)))). (-32.98 = -32.66 + -0.32)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:02 2006