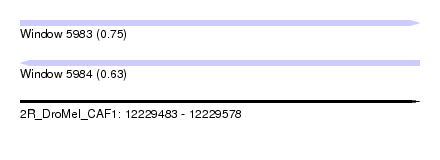

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,229,483 – 12,229,578 |
| Length | 95 |
| Max. P | 0.748755 |

| Location | 12,229,483 – 12,229,578 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.25 |
| Mean single sequence MFE | -23.54 |
| Consensus MFE | -15.51 |
| Energy contribution | -15.32 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748755 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12229483 95 + 20766785 AAAUUGCUUAAGCUUGCUCAAAUCCGAUUUAACUAGCUAGCAUGGGCACACUGUGAUUUUAGUGUGGCCUACUGAUUAUAAUAAGC-------------------UACCUAUAA--- .....(((((.(((.(((................))).)))..((.(((((((......))))))).))............)))))-------------------.........--- ( -23.69) >DroSec_CAF1 90545 95 + 1 AAAUUGCUUCAGCUUGCUCAAAUCCGAUUUAACUAGCUAGCAUGGGCACACUGUGAUUUUAGUGUGGCCUACUGAUUAUAGCAAGC-------------------UACCUAUAA--- ..........((((((((..((((.(.........(....)..((.(((((((......))))))).))..).))))..)))))))-------------------)........--- ( -27.20) >DroSim_CAF1 86997 95 + 1 AAAUUGCUUCAGCUUGCUCAAAUCCGAUUGAACUAGCUAGCAUGGGCACAUUAUGAUUUUAGUGUGGACUACUGAUUAUAACAAGU-------------------UACCUAUAA--- .....(((..((((.(.((((......)))).).)))))))((((((((((((......)))))))((((.............)))-------------------).)))))..--- ( -20.22) >DroYak_CAF1 93536 117 + 1 GAAUUGAUUCAGCUUGCCCUAAGCCCAUUUAACAAGCUAGCAAGGUCACACUGUGCUUUUAGUGUGGUCUACUGAUUAUAAUAAGCGCGCCCAAUAAUUACACGCUCCCAACAAAUU ...(((....((((((................))))))(((.((..(((((((......)))))))..))..(((((((..............)))))))...))).....)))... ( -23.03) >consensus AAAUUGCUUCAGCUUGCUCAAAUCCGAUUUAACUAGCUAGCAUGGGCACACUGUGAUUUUAGUGUGGCCUACUGAUUAUAACAAGC___________________UACCUAUAA___ .....((((..(((.((..................)).)))..((.(((((((......))))))).)).............))))............................... (-15.51 = -15.32 + -0.19)


| Location | 12,229,483 – 12,229,578 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 79.25 |
| Mean single sequence MFE | -25.90 |
| Consensus MFE | -15.10 |
| Energy contribution | -15.98 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.629698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12229483 95 - 20766785 ---UUAUAGGUA-------------------GCUUAUUAUAAUCAGUAGGCCACACUAAAAUCACAGUGUGCCCAUGCUAGCUAGUUAAAUCGGAUUUGAGCAAGCUUAAGCAAUUU ---.........-------------------(((((........((((((.((((((........)))))).)).))))((((.(((.((......)).))).)))))))))..... ( -22.30) >DroSec_CAF1 90545 95 - 1 ---UUAUAGGUA-------------------GCUUGCUAUAAUCAGUAGGCCACACUAAAAUCACAGUGUGCCCAUGCUAGCUAGUUAAAUCGGAUUUGAGCAAGCUGAAGCAAUUU ---.......((-------------------(((((((..((((((((((.((((((........)))))).)).)))).....(......).))))..)))))))))......... ( -27.10) >DroSim_CAF1 86997 95 - 1 ---UUAUAGGUA-------------------ACUUGUUAUAAUCAGUAGUCCACACUAAAAUCAUAAUGUGCCCAUGCUAGCUAGUUCAAUCGGAUUUGAGCAAGCUGAAGCAAUUU ---.....((((-------------------....(((((.((...((((....))))..)).))))).))))..((((((((.((((((......)))))).))))..)))).... ( -20.90) >DroYak_CAF1 93536 117 - 1 AAUUUGUUGGGAGCGUGUAAUUAUUGGGCGCGCUUAUUAUAAUCAGUAGACCACACUAAAAGCACAGUGUGACCUUGCUAGCUUGUUAAAUGGGCUUAGGGCAAGCUGAAUCAAUUC .....(((((((((((((.........)))))))).......((((.....((((((........))))))..((((((((((..(...)..))))...)))))))))).))))).. ( -33.30) >consensus ___UUAUAGGUA___________________GCUUAUUAUAAUCAGUAGGCCACACUAAAAUCACAGUGUGCCCAUGCUAGCUAGUUAAAUCGGAUUUGAGCAAGCUGAAGCAAUUU ........((.....................(((((((......)))))))((((((........)))))).)).((((((((.(((.((......)).))).))))..)))).... (-15.10 = -15.98 + 0.87)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:44 2006