
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,153,492 – 12,153,583 |
| Length | 91 |
| Max. P | 0.757811 |

| Location | 12,153,492 – 12,153,583 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 90.50 |
| Mean single sequence MFE | -32.16 |
| Consensus MFE | -25.42 |
| Energy contribution | -25.98 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.757811 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
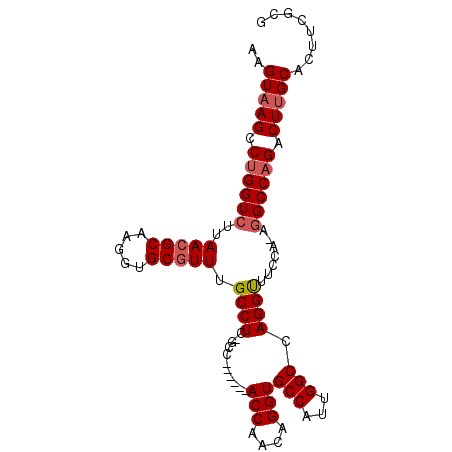
>2R_DroMel_CAF1 12153492 91 + 20766785 AAGUAAGCCUGGUCUUAACGCAAGGUGCAUUUGCCUC-GCC------ACCAACAGGUGCCAUUGGCCAGGUUUGCAAAGGCCCGACUUGCACUUCGCG ..((((((((((((.....((.((((......)))).-))(------(((....)))).....)))))))))))).(((......)))((.....)). ( -33.10) >DroSec_CAF1 9548 91 + 1 AAGUAAGCCUGGUCUUAACGCAAGGUGCGUUUGCCUC-GCC------ACCAACAGGUGCCAUUGGCCAGGUUUCCAACGGCCAGACUUGCACUUCGCG ..(((((.((((((..(((((.....))))).((((.-(((------(((....)))......))).)))).......)))))).)))))........ ( -31.10) >DroSim_CAF1 9633 91 + 1 AAGUAAGCCUGGUCUUAACGCAAGGUGCGUUUGCCUC-GCC------ACCAACAGGUGCCAUUGGCCAGGUUUCCAGCGGCCAGACUUGCACUUCGCG ..(((((.((((((....(....)..((....((((.-(((------(((....)))......))).)))).....)))))))).)))))........ ( -31.10) >DroEre_CAF1 15227 90 + 1 AAGUAAGCCUGGUCUUAACGCAAGGUGCGUUUGCCUC-GAC------ACCAACAGGUGCCAUUGGCAAGGCUUUAA-AGGCCAGACUCGCACUUCGCG ..(..((.((((((((..(....)..((.((((((..-(.(------(((....)))).)...))))))))....)-))))))).))..)........ ( -31.60) >DroYak_CAF1 15384 97 + 1 AAGUAAGCCUGGUCUUAACGCAAGGUGCGUUUGCCUCCGACAGGCAAACCAACAGGUGCCAUUGGCCAGGCUUUUA-AAGCCAGACUUGCACUUCGCG ....((((((((((....(((....((.((((((((.....))))))))))....))).....))))))))))...-...........((.....)). ( -33.90) >consensus AAGUAAGCCUGGUCUUAACGCAAGGUGCGUUUGCCUC_GCC______ACCAACAGGUGCCAUUGGCCAGGUUUCCA_AGGCCAGACUUGCACUUCGCG ..(((((.((((((..(((((.....))))).((((...........(((....)))(((...))).)))).......)))))).)))))........ (-25.42 = -25.98 + 0.56)


| Location | 12,153,492 – 12,153,583 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 90.50 |
| Mean single sequence MFE | -34.16 |
| Consensus MFE | -25.57 |
| Energy contribution | -26.01 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.648915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
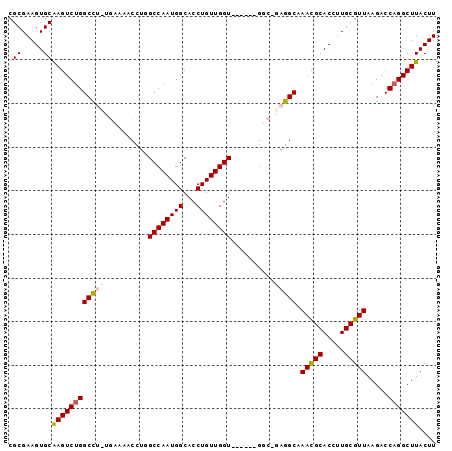
>2R_DroMel_CAF1 12153492 91 - 20766785 CGCGAAGUGCAAGUCGGGCCUUUGCAAACCUGGCCAAUGGCACCUGUUGGU------GGC-GAGGCAAAUGCACCUUGCGUUAAGACCAGGCUUACUU .((((((.((.......))))))))...(((((.(.....((((....)))------)((-((((........)))))).....).)))))....... ( -32.30) >DroSec_CAF1 9548 91 - 1 CGCGAAGUGCAAGUCUGGCCGUUGGAAACCUGGCCAAUGGCACCUGUUGGU------GGC-GAGGCAAACGCACCUUGCGUUAAGACCAGGCUUACUU .((.....))(((((((((((((((........)))))))((((....)))------)((-((((........)))))).......)))))))).... ( -34.70) >DroSim_CAF1 9633 91 - 1 CGCGAAGUGCAAGUCUGGCCGCUGGAAACCUGGCCAAUGGCACCUGUUGGU------GGC-GAGGCAAACGCACCUUGCGUUAAGACCAGGCUUACUU .((.....))((((((((((((..(....)..))...((.((((....)))------).)-).))).(((((.....))))).....))))))).... ( -33.60) >DroEre_CAF1 15227 90 - 1 CGCGAAGUGCGAGUCUGGCCU-UUAAAGCCUUGCCAAUGGCACCUGUUGGU------GUC-GAGGCAAACGCACCUUGCGUUAAGACCAGGCUUACUU .((.....))((((((((.((-(........((((..(((((((....)))------)))-).))))(((((.....)))))))).)))))))).... ( -35.20) >DroYak_CAF1 15384 97 - 1 CGCGAAGUGCAAGUCUGGCUU-UAAAAGCCUGGCCAAUGGCACCUGUUGGUUUGCCUGUCGGAGGCAAACGCACCUUGCGUUAAGACCAGGCUUACUU (((((.((((..(((.((((.-....)))).)))((((((...))))))((((((((.....)))))))))))).))))).((((......))))... ( -35.00) >consensus CGCGAAGUGCAAGUCUGGCCU_UGAAAACCUGGCCAAUGGCACCUGUUGGU______GGC_GAGGCAAACGCACCUUGCGUUAAGACCAGGCUUACUU .((.....))((((((((((((..........((((((((...))))))))..........))))).(((((.....))))).....))))))).... (-25.57 = -26.01 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:11 2006