
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 12,092,386 – 12,092,476 |
| Length | 90 |
| Max. P | 0.961804 |

| Location | 12,092,386 – 12,092,476 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 89.02 |
| Mean single sequence MFE | -32.46 |
| Consensus MFE | -27.34 |
| Energy contribution | -28.38 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.572706 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12092386 90 + 20766785 AUUCGCUGGGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGUUUGUCGAUUUCAAGAC------------CCGUCUUGGCCGUGUCAAUGGCCAACCGGUUGC ..(((...(((((.(((((....(((((((((((.........(((........)))------------))).))))))))))))).)))))...))).... ( -33.50) >DroSec_CAF1 5773 90 + 1 AUUCGUUGGGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGUUUGUCGAUUUCGAGAC------------CCGUCUUGGCCGUGUCAAUGGCCAACCGGUUGC ..(((...(((((.(((((....(((((((((((.....(((..(((....))))))------------))).))))))))))))).)))))...))).... ( -34.10) >DroSim_CAF1 4911 90 + 1 AUUCGUUGGGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGUUUGUCGAUUUCGAGAC------------CCGUCUUGGCCGUGUCAAUGGCCAACCGGUUGC ..(((...(((((.(((((....(((((((((((.....(((..(((....))))))------------))).))))))))))))).)))))...))).... ( -34.10) >DroEre_CAF1 5852 89 + 1 GUUCGAAGAGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGCUU-UCGAUUUCAAGAC------------CCGUCUUGGCCGUGUCAAUGGCCAACCGGUUGC ..((((.((((.(((..((....))(((....)))))).))))-))))......(((------------(.((..(((((((....))))))))).)))).. ( -27.20) >DroYak_CAF1 6348 101 + 1 AUGCGAAGAGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGUUU-UCGAUUUCAAGACUCGGAUUUGGACUCGUCGCGGCCGUGUCAAUGGCCAACCGGUUGC ..((((.(((((((...((......(((....)))...(((((-(.(....)))))))))...)))).))).))))((((((....)))))).......... ( -33.40) >consensus AUUCGAUGGGCCAGUGACGUUGUUGGCCAAGUGGCAUUAGUUUGUCGAUUUCAAGAC____________CCGUCUUGGCCGUGUCAAUGGCCAACCGGUUGC ...((...(((((.(((((....(((((((((((.........(((........)))............))).))))))))))))).)))))...))..... (-27.34 = -28.38 + 1.04)
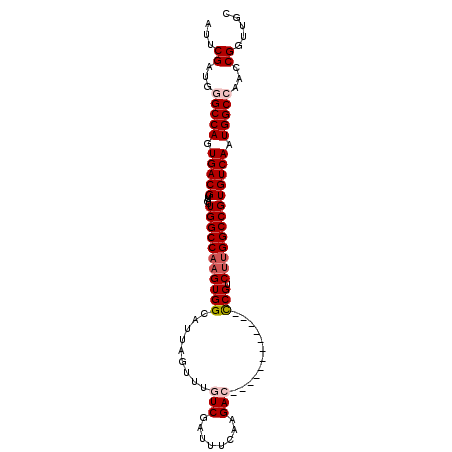



| Location | 12,092,386 – 12,092,476 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 89.02 |
| Mean single sequence MFE | -29.24 |
| Consensus MFE | -23.62 |
| Energy contribution | -23.22 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 12092386 90 - 20766785 GCAACCGGUUGGCCAUUGACACGGCCAAGACGG------------GUCUUGAAAUCGACAAACUAAUGCCACUUGGCCAACAACGUCACUGGCCCAGCGAAU .....((.(.(((((.((((..(((((((...(------------((.(((............))).))).)))))))......)))).))))).).))... ( -29.80) >DroSec_CAF1 5773 90 - 1 GCAACCGGUUGGCCAUUGACACGGCCAAGACGG------------GUCUCGAAAUCGACAAACUAAUGCCACUUGGCCAACAACGUCACUGGCCCAACGAAU .......((((((((.((((..(((((((...(------------((.(((....))).........))).)))))))......)))).))).))))).... ( -29.60) >DroSim_CAF1 4911 90 - 1 GCAACCGGUUGGCCAUUGACACGGCCAAGACGG------------GUCUCGAAAUCGACAAACUAAUGCCACUUGGCCAACAACGUCACUGGCCCAACGAAU .......((((((((.((((..(((((((...(------------((.(((....))).........))).)))))))......)))).))).))))).... ( -29.60) >DroEre_CAF1 5852 89 - 1 GCAACCGGUUGGCCAUUGACACGGCCAAGACGG------------GUCUUGAAAUCGA-AAGCUAAUGCCACUUGGCCAACAACGUCACUGGCUCUUCGAAC .....(((..(((((.((((..(((((((...(------------((.(((....(..-..).))).))).)))))))......)))).)))))..)))... ( -30.60) >DroYak_CAF1 6348 101 - 1 GCAACCGGUUGGCCAUUGACACGGCCGCGACGAGUCCAAAUCCGAGUCUUGAAAUCGA-AAACUAAUGCCACUUGGCCAACAACGUCACUGGCUCUUCGCAU ..........((((........))))((((.(((.(((....(((((.(((....)))-..)))...(((....)))......))....)))))).)))).. ( -26.60) >consensus GCAACCGGUUGGCCAUUGACACGGCCAAGACGG____________GUCUUGAAAUCGACAAACUAAUGCCACUUGGCCAACAACGUCACUGGCCCAACGAAU ....((((((((((........))))).((((................(((....))).........(((....)))......))))))))).......... (-23.62 = -23.22 + -0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:41 2006