
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,793,744 – 11,793,834 |
| Length | 90 |
| Max. P | 0.999210 |

| Location | 11,793,744 – 11,793,834 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 77.99 |
| Mean single sequence MFE | -27.20 |
| Consensus MFE | -21.88 |
| Energy contribution | -22.63 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.80 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986882 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
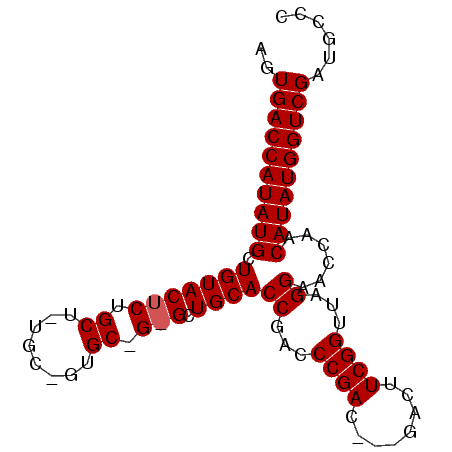
>2R_DroMel_CAF1 11793744 90 + 20766785 AGUGACCAUAUGCUGUACUCUGCUGUGCCGUGCUGAGCUGCACCGACCCGACUUUGACUUCGGUUAGGAACCAACAUAUGGUCGAUGCCC ..((((((((((.(((((((.((........)).))).))))((...((((........))))...))......))))))))))...... ( -28.50) >DroSim_CAF1 15106 84 + 1 AGUGACCAUAUGCUGUACUCUGCUCUGCUGUGCCGUGCUGCACCGACCCGAC------UUCGGUUAGGAACCAACAUAUGGUCGAUGCCC ..(((((((((((.((((...((.(....).)).)))).)).....((..((------....))..)).......)))))))))...... ( -26.30) >DroEre_CAF1 14202 68 + 1 AGUGACCAUAUGUUGUA--------------------CUGCACCGACCCGAC--CGACUUCGGUUAGGAACCAACAUAUGGUCGAUGCCC ..(((((((((((((..--------------------(......).((..((--((....))))..))...)))))))))))))...... ( -26.80) >consensus AGUGACCAUAUGCUGUACUCUGCU_UGC_GUGC_G_GCUGCACCGACCCGAC___GACUUCGGUUAGGAACCAACAUAUGGUCGAUGCCC ..((((((((((.(((((((.((........)).))).))))((...((((........))))...))......))))))))))...... (-21.88 = -22.63 + 0.75)


| Location | 11,793,744 – 11,793,834 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 77.99 |
| Mean single sequence MFE | -30.48 |
| Consensus MFE | -28.04 |
| Energy contribution | -28.57 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.44 |
| SVM RNA-class probability | 0.999210 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
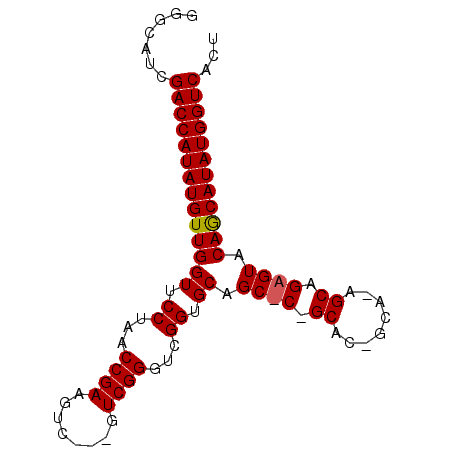
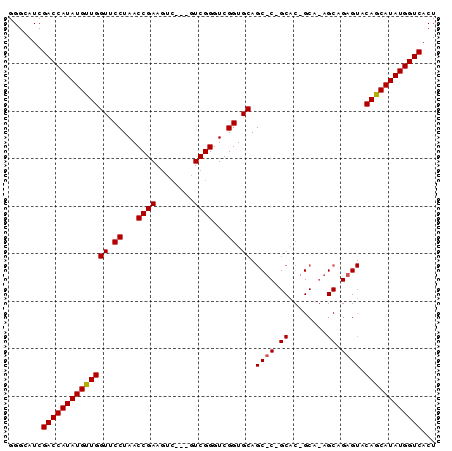
>2R_DroMel_CAF1 11793744 90 - 20766785 GGGCAUCGACCAUAUGUUGGUUCCUAACCGAAGUCAAAGUCGGGUCGGUGCAGCUCAGCACGGCACAGCAGAGUACAGCAUAUGGUCACU .......((((((((((((((.((...((((........))))...)).)).((((.((........)).)))).))))))))))))... ( -34.00) >DroSim_CAF1 15106 84 - 1 GGGCAUCGACCAUAUGUUGGUUCCUAACCGAA------GUCGGGUCGGUGCAGCACGGCACAGCAGAGCAGAGUACAGCAUAUGGUCACU .......((((((((((((((.((...(((..------..)))...)).)).((.(.((...)).).))......))))))))))))... ( -29.90) >DroEre_CAF1 14202 68 - 1 GGGCAUCGACCAUAUGUUGGUUCCUAACCGAAGUCG--GUCGGGUCGGUGCAG--------------------UACAACAUAUGGUCACU .......((((((((((((...((..((((....))--))..)).........--------------------..))))))))))))... ( -27.54) >consensus GGGCAUCGACCAUAUGUUGGUUCCUAACCGAAGUC___GUCGGGUCGGUGCAGC_C_GCAC_GCA_AGCAGAGUACAGCAUAUGGUCACU .......((((((((((((((.((...((((........))))...)).)).((((.((........)).)))).))))))))))))... (-28.04 = -28.57 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:32 2006