
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,620,846 – 11,620,953 |
| Length | 107 |
| Max. P | 0.736121 |

| Location | 11,620,846 – 11,620,953 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 80.19 |
| Mean single sequence MFE | -21.27 |
| Consensus MFE | -13.88 |
| Energy contribution | -14.38 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.736121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
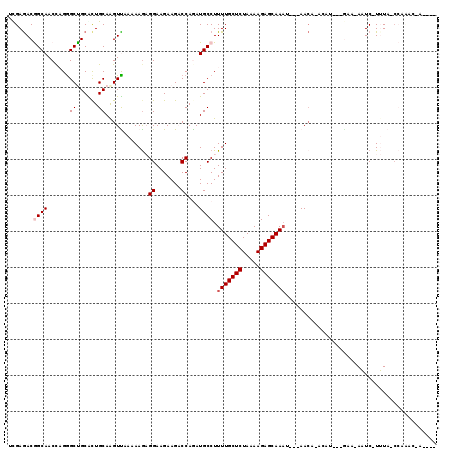
>2R_DroMel_CAF1 11620846 107 + 20766785 UCGAGACGGCAACUAGGGCCGCGCUGCAAGUCAAAAAGAGGAAGAAGACCAGAUGCCUUUUGCUCUAAAAGAGCAAAU--CAACA-ACAU---GAA-AAUC-UUUA-CCACAC-ACA-- ....((((((.......)))((...))..))).......((..(((((..........(((((((.....)))))))(--((...-...)---)).-..))-))).-))....-...-- ( -20.30) >DroGri_CAF1 2232 105 + 1 UCGAGACAGCAACCAGGGCUGCACUGCAAGUUAAAAAGCGGAAGAAGCCCAAAUGCCUUUUGCUCUAAAAGAGCAAAU---AACA-AAAA-A-AAGAAUUC-UAUA-C-AUAC-A---- ...(((..(((....(((((...((((..........))))....)))))...)))..(((((((.....))))))).---....-....-.-......))-)...-.-....-.---- ( -23.70) >DroWil_CAF1 9322 109 + 1 UUGAAACGGCAACCCGAGCUGCACUGCAAGUUAAAAAGAGGAAGAAGACCAGAUGCCUUUUGCUCUAAAAGAGCAAAAUACAAAA-ACACAAAGCA-AAUCGUUUA-ACAAA------- ...((((((((......((......))............((.......))...))))((((((((.....)))))))).......-..........-....)))).-.....------- ( -19.60) >DroMoj_CAF1 1805 98 + 1 UCGAGACAGCAACCAGGGCUGCACUGCAAGUUAAAAAACGGAAGAAGCCCAAAUGCCUUUUGCUCUAAAAGAGCAAAU---AACA-AA---------AUUC-UAUA-C-AAAC-A---- ...(((..(((....(((((...((....(((....)))...)).)))))...)))..(((((((.....))))))).---....-..---------..))-)...-.-....-.---- ( -20.70) >DroAna_CAF1 2045 103 + 1 UUGAGACGGCAACCCGGGCCGCGCUGCAAGUCAAAAAGAGGAAGAAGACCAGAUGCCUUUUGCUCUAAAAGAGCAAAU--CAACAAACAU---GAA-AAUC-UUUA-CCAC-------- ....((((((.......)))((...))..))).......((..(((((..........(((((((.....)))))))(--((.......)---)).-..))-))).-))..-------- ( -19.80) >DroPer_CAF1 4221 109 + 1 UUGAGACGGCAACCAGGGCUGCGUUGCAGGUUAAAAAGAGGAAGAAGACCAGAUGCCUUUUGCUCUAAAAGAGCAACU---AACA-A-AU---GAA-AAUC-UUUAACCAUAAUACAAC ........(((((((....)).))))).(((((((((((((..............))))))((((.....))))....---....-.-..---...-....-))))))).......... ( -23.54) >consensus UCGAGACGGCAACCAGGGCUGCACUGCAAGUUAAAAAGAGGAAGAAGACCAGAUGCCUUUUGCUCUAAAAGAGCAAAU___AACA_ACAU___GAA_AAUC_UUUA_CCAAAC_A____ .......((((......((......))............((.......))...)))).(((((((.....))))))).......................................... (-13.88 = -14.38 + 0.50)


| Location | 11,620,846 – 11,620,953 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 80.19 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -17.61 |
| Energy contribution | -17.42 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.709871 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
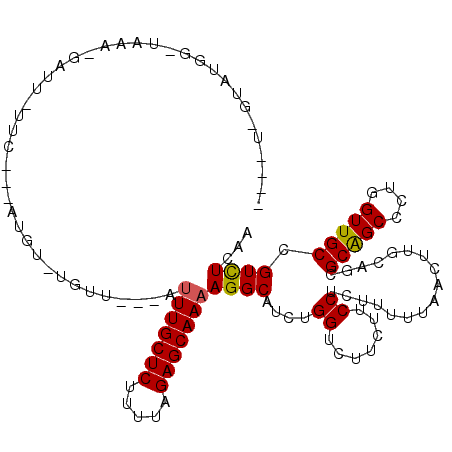
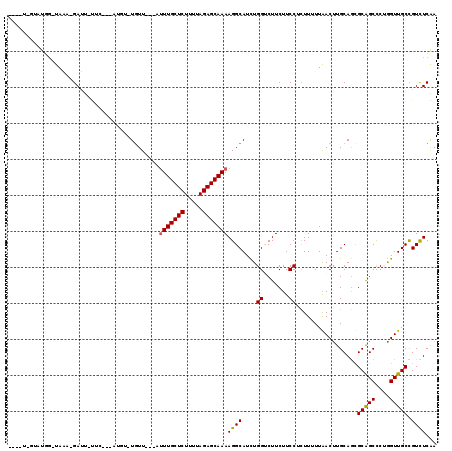
>2R_DroMel_CAF1 11620846 107 - 20766785 --UGU-GUGUGG-UAAA-GAUU-UUC---AUGU-UGUUG--AUUUGCUCUUUUAGAGCAAAAGGCAUCUGGUCUUCUUCCUCUUUUUGACUUGCAGCGCGGCCCUAGUUGCCGUCUCGA --...-..(.((-..((-((((-...---((((-(....--.(((((((.....))))))).)))))..))))))...)).).....(((..(((((.........))))).))).... ( -24.60) >DroGri_CAF1 2232 105 - 1 ----U-GUAU-G-UAUA-GAAUUCUU-U-UUUU-UGUU---AUUUGCUCUUUUAGAGCAAAAGGCAUUUGGGCUUCUUCCGCUUUUUAACUUGCAGUGCAGCCCUGGUUGCUGUCUCGA ----(-(((.-(-(.((-(((..(..-.-....-(((.---.(((((((.....)))))))..)))...(((.....))))..))))))).))))..(((((....)))))........ ( -24.30) >DroWil_CAF1 9322 109 - 1 -------UUUGU-UAAACGAUU-UGCUUUGUGU-UUUUGUAUUUUGCUCUUUUAGAGCAAAAGGCAUCUGGUCUUCUUCCUCUUUUUAACUUGCAGUGCAGCUCGGGUUGCCGUUUCAA -------.....-.(((((..(-(((...((..-...(((.((((((((.....)))))))).)))...((.......))........))..)))).(((((....))))))))))... ( -24.30) >DroMoj_CAF1 1805 98 - 1 ----U-GUUU-G-UAUA-GAAU---------UU-UGUU---AUUUGCUCUUUUAGAGCAAAAGGCAUUUGGGCUUCUUCCGUUUUUUAACUUGCAGUGCAGCCCUGGUUGCUGUCUCGA ----(-((..-(-(.((-(((.---------..-(((.---.(((((((.....)))))))..)))...(((.....)))...)))))))..)))..(((((....)))))........ ( -23.70) >DroAna_CAF1 2045 103 - 1 --------GUGG-UAAA-GAUU-UUC---AUGUUUGUUG--AUUUGCUCUUUUAGAGCAAAAGGCAUCUGGUCUUCUUCCUCUUUUUGACUUGCAGCGCGGCCCGGGUUGCCGUCUCAA --------(.((-..((-((((-...---((((((....--.(((((((.....)))))))))))))..))))))...)).).....(((..(((((.((...)).))))).))).... ( -27.10) >DroPer_CAF1 4221 109 - 1 GUUGUAUUAUGGUUAAA-GAUU-UUC---AU-U-UGUU---AGUUGCUCUUUUAGAGCAAAAGGCAUCUGGUCUUCUUCCUCUUUUUAACCUGCAACGCAGCCCUGGUUGCCGUCUCAA ((((((....(((((((-((..-.((---(.-.-((((---..((((((.....))))))..))))..)))...)))........))))))))))))(((((....)))))........ ( -31.50) >consensus ____U_GUAUGG_UAAA_GAUU_UUC___AUGU_UGUU___AUUUGCUCUUUUAGAGCAAAAGGCAUCUGGUCUUCUUCCUCUUUUUAACUUGCAGCGCAGCCCUGGUUGCCGUCUCAA ..........................................(((((((.....)))))))((((....((.......)).................(((((....))))).))))... (-17.61 = -17.42 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:03 2006