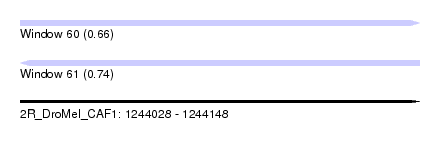
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 1,244,028 – 1,244,148 |
| Length | 120 |
| Max. P | 0.740985 |

| Location | 1,244,028 – 1,244,148 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -37.71 |
| Consensus MFE | -35.24 |
| Energy contribution | -35.05 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.661264 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 1244028 120 + 20766785 UUAUUUGUCAUGACUUUUUCCCUUAUCAGCCGAGCCUCGAGUAGCUGUCCCUGGAGUAGUCGCAACGCCGCUUGAAGGGUGAAGCUACAGAGUUCACCCUCCCAGUGGCAAGCUUUGACA .....(((((.((((..((((.....((((((.....))....)))).....)))).))))((...((((((.(.((((((((.((....)))))))))).).))))))..))..))))) ( -42.20) >DroSim_CAF1 253 119 + 1 UUAUUUGUCAUGACUUUUUCCCUUAUCAGCCGAGCCUCGAGUAGCUGUCCCUGGAGUGGUCCCAACGCCGCUUGAAGGGUGAAGCU-CAGAGUUCACCCUCCCGGUGGCAAGCUUUGACA .....(((((.((((..((((.....((((((.....))....)))).....)))).)))).....((((((.(.((((((((...-.....)))))))).).))))))......))))) ( -39.80) >DroEre_CAF1 63 120 + 1 UUAUUUGUCAUGACUUUUUCCCUUAUCAGCCGAACCUCGAGUUGUUGUCCCUGGAGUAGUCCCAAAGGCGCUUGAGGGGUGAAGCUACAGUGUCCACCCUCCCAGUGGCAAGCUUUGACA .....(((((.((((..((((.....((((((.....)).))))........)))).)))).....(.((((.(..(((((..((....))...)))))..).)))).)......))))) ( -33.42) >DroYak_CAF1 258 120 + 1 UUAUUUGUCAUGACUUUUUCCCUUAUCAGCCGAACCUUGAAUAGCUGUCCCUGGAGUAGUCCCAAAGCCGCUUGAAGGGUGAAGCUACAGUGUUCACCCUCCCAGUGGAAAGCUUUGACA .....(((((.((((..((((.....((((.............)))).....)))).))))......(((((.(.((((((((((....)).)))))))).).))))).......))))) ( -35.42) >consensus UUAUUUGUCAUGACUUUUUCCCUUAUCAGCCGAACCUCGAGUAGCUGUCCCUGGAGUAGUCCCAAAGCCGCUUGAAGGGUGAAGCUACAGAGUUCACCCUCCCAGUGGCAAGCUUUGACA .....(((((.((((..((((.....((((((.....))....)))).....)))).)))).....((((((.(.(((((((.((......))))))))).).))))))......))))) (-35.24 = -35.05 + -0.19)

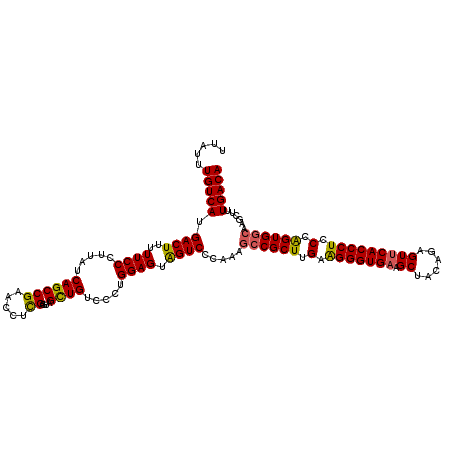
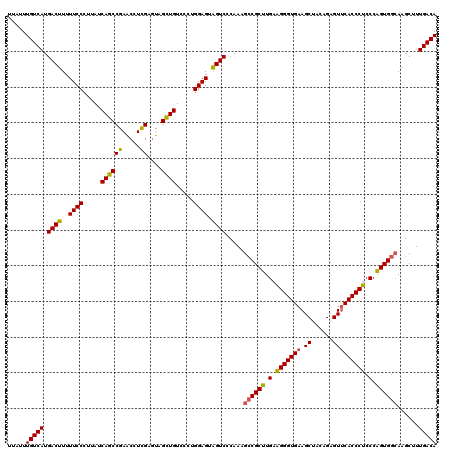
| Location | 1,244,028 – 1,244,148 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -39.78 |
| Consensus MFE | -33.16 |
| Energy contribution | -33.73 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740985 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 1244028 120 - 20766785 UGUCAAAGCUUGCCACUGGGAGGGUGAACUCUGUAGCUUCACCCUUCAAGCGGCGUUGCGACUACUCCAGGGACAGCUACUCGAGGCUCGGCUGAUAAGGGAAAAAGUCAUGACAAAUAA (((((..((.((((.((.((((((((((((....)).)))))))))).)).))))..))((((..(((.....(((((...........))))).....)))...)))).)))))..... ( -44.60) >DroSim_CAF1 253 119 - 1 UGUCAAAGCUUGCCACCGGGAGGGUGAACUCUG-AGCUUCACCCUUCAAGCGGCGUUGGGACCACUCCAGGGACAGCUACUCGAGGCUCGGCUGAUAAGGGAAAAAGUCAUGACAAAUAA (((((..(((((((.(..((((((((((.....-...))))))))))..).)))((((((....(((.((.........)).))).))))))............))))..)))))..... ( -40.40) >DroEre_CAF1 63 120 - 1 UGUCAAAGCUUGCCACUGGGAGGGUGGACACUGUAGCUUCACCCCUCAAGCGCCUUUGGGACUACUCCAGGGACAACAACUCGAGGUUCGGCUGAUAAGGGAAAAAGUCAUGACAAAUAA (((((..((((.((..((((.((((((((......).)))))))))))...(((.(((((.....)))))((((..(.....)..))))))).......))...))))..)))))..... ( -35.90) >DroYak_CAF1 258 120 - 1 UGUCAAAGCUUUCCACUGGGAGGGUGAACACUGUAGCUUCACCCUUCAAGCGGCUUUGGGACUACUCCAGGGACAGCUAUUCAAGGUUCGGCUGAUAAGGGAAAAAGUCAUGACAAAUAA (((((..(((((((.((.(((((((((((......).)))))))))).)).))(((((((.....))))))).(((((...........))))).........)))))..)))))..... ( -38.20) >consensus UGUCAAAGCUUGCCACUGGGAGGGUGAACACUGUAGCUUCACCCUUCAAGCGGCGUUGGGACUACUCCAGGGACAGCUACUCGAGGCUCGGCUGAUAAGGGAAAAAGUCAUGACAAAUAA (((((..((((.((....(((((((((((......).)))))))))).((((((((((((.....((....))......)))))))))..)))......))...))))..)))))..... (-33.16 = -33.73 + 0.56)

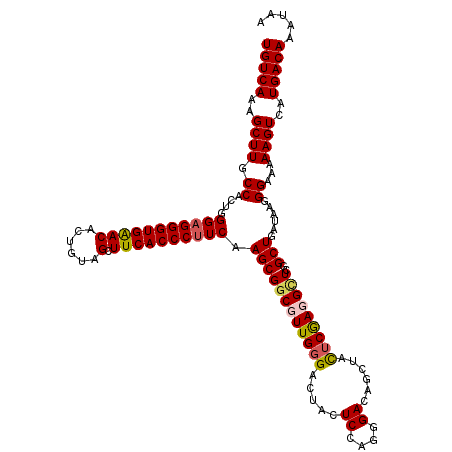
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:24:05 2006