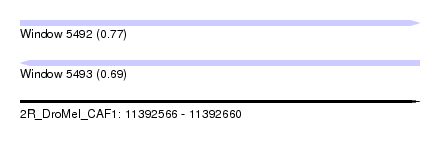

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,392,566 – 11,392,660 |
| Length | 94 |
| Max. P | 0.765703 |

| Location | 11,392,566 – 11,392,660 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 84.35 |
| Mean single sequence MFE | -28.03 |
| Consensus MFE | -17.74 |
| Energy contribution | -16.90 |
| Covariance contribution | -0.84 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.765703 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11392566 94 + 20766785 UCCUUCAGCUUUCCCUUCCCCUCCGACCACUUUCC--CUGGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCCUUAGGAGCGGAGUA ..((((.(((((.....((((((((.........(--(((((........)))))).............))))...)))).....))))))))).. ( -27.95) >DroSec_CAF1 5658 85 + 1 UCCUUCAGAUCCCCUUUC---------CACUUUCC--CUGGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCUUUAGGAGCGGAGUA ((((..((..(((((.((---------(.......--(((((........)))))(((.........))))))..)))))))..))))........ ( -25.50) >DroSim_CAF1 5254 85 + 1 UCCUUUAGCUCCCCUUUC---------CACUUUCC--CUGGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCUUUAGGAGCGGAGUA ((((..((((((....((---------(.......--(((((........)))))(((.........))))))...))))))..))))........ ( -27.70) >DroEre_CAF1 8168 91 + 1 UCCCUCAGCUUCCU---CAGCGCCCUCCGCAUUCC--CUGGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCUUUAGGAGCGGAGUA ...(((.(((.(((---.(((.((((((((((..(--(((((........))))))..........)))))))...))))))..)))))).))).. ( -37.10) >DroYak_CAF1 6189 96 + 1 UCCCUCAGCUCCCUUCACCACUCCCUCCAUCUUCCAACUCGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCUUUAGGAGCGGAGUA .......(((((((((.....(((((......(((..((.((........)).))(((.........))))))..))))).....)))).))))). ( -21.90) >consensus UCCUUCAGCUCCCCUUUC__C_CC__CCACUUUCC__CUGGCCAAUCCAUGCCAGGCACUUUAAUUAUGCGGACCAGGGGCUUUAGGAGCGGAGUA ((((((....((((.......................(((((........)))))(((.........)))......))))......))).)))... (-17.74 = -16.90 + -0.84)

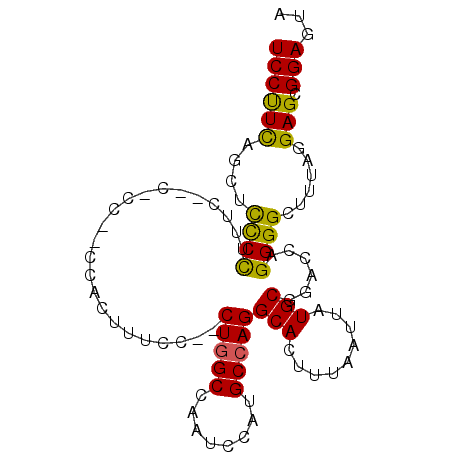
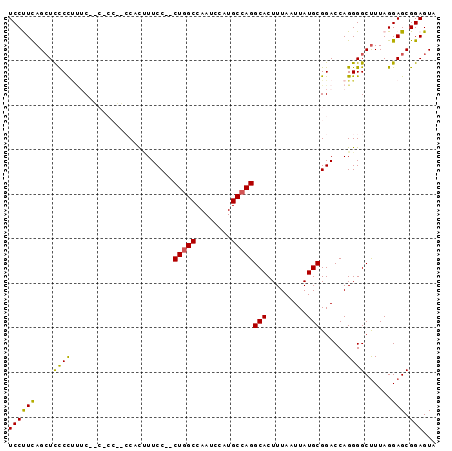
| Location | 11,392,566 – 11,392,660 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 84.35 |
| Mean single sequence MFE | -33.70 |
| Consensus MFE | -19.87 |
| Energy contribution | -20.23 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11392566 94 - 20766785 UACUCCGCUCCUAAGGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCCAG--GGAAAGUGGUCGGAGGGGAAGGGAAAGCUGAAGGA ..(((..(((((..((((.((..(((((((.......))))(((((........)))))--))).)).))))..)))))..)))............ ( -34.20) >DroSec_CAF1 5658 85 - 1 UACUCCGCUCCUAAAGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCCAG--GGAAAGUG---------GAAAGGGGAUCUGAAGGA ........((((..(((((((..(((((..........(.((((((........)))))--).)..)))---------)).)))))..))..)))) ( -30.40) >DroSim_CAF1 5254 85 - 1 UACUCCGCUCCUAAAGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCCAG--GGAAAGUG---------GAAAGGGGAGCUAAAGGA ........((((..(((((((..(((((..........(.((((((........)))))--).)..)))---------))..)))).)))..)))) ( -33.10) >DroEre_CAF1 8168 91 - 1 UACUCCGCUCCUAAAGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCCAG--GGAAUGCGGAGGGCGCUG---AGGAAGCUGAGGGA ..(((.((((((..((((((...(((((((........(.((((((........)))))--).))))))))))).))).---))).))).)))... ( -40.10) >DroYak_CAF1 6189 96 - 1 UACUCCGCUCCUAAAGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCGAGUUGGAAGAUGGAGGGAGUGGUGAAGGGAGCUGAGGGA ..(((.((((((...(((.((..(((.(((..((((..((((...........))))..))))...)))..))))).)))...)))))).)))... ( -30.70) >consensus UACUCCGCUCCUAAAGCCCCUGGUCCGCAUAAUUAAAGUGCCUGGCAUGGAUUGGCCAG__GGAAAGUGG__GG_G__GAAAGGGGAGCUGAAGGA ........((((..(((((((..(((((((.......))))(((((........)))))..)))..................)))).)))..)))) (-19.87 = -20.23 + 0.36)

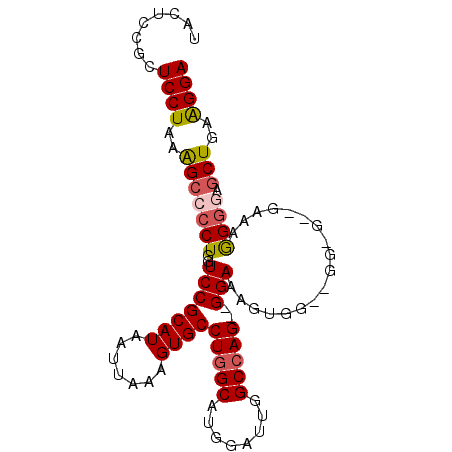
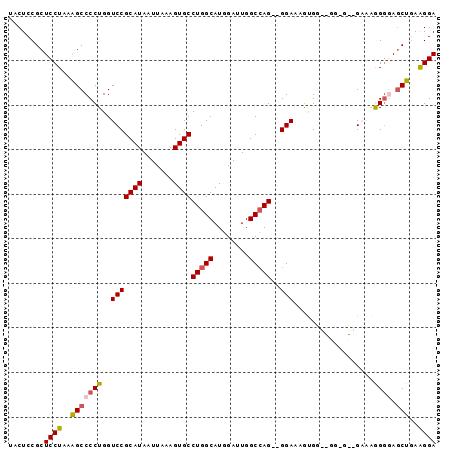
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:38 2006