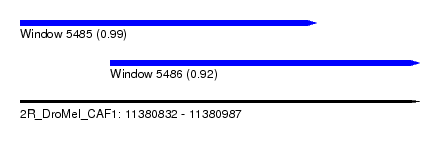

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,380,832 – 11,380,987 |
| Length | 155 |
| Max. P | 0.990442 |

| Location | 11,380,832 – 11,380,947 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.51 |
| Mean single sequence MFE | -43.29 |
| Consensus MFE | -36.73 |
| Energy contribution | -35.66 |
| Covariance contribution | -1.06 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.87 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.990442 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
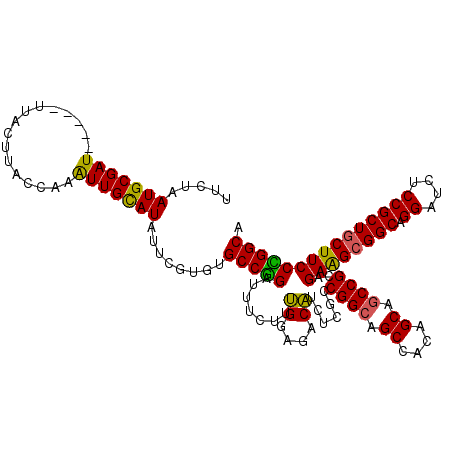
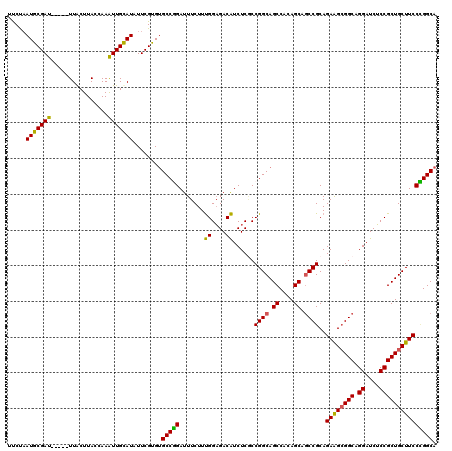
>2R_DroMel_CAF1 11380832 115 + 20766785 GUCUAAUGCGAU-----UUACUUACCAAAUUGUAUAUUCGCGUGCCUGAUUUCUUUGGAGACAUCUCACCGGCAGCCACAGCCCCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCAGGCA .....((((((.-----.(((..........)))...))))))(((((.........(((....)))..(((..((....))..)))..((((((((.((....))))))))))))))). ( -39.20) >DroSec_CAF1 8093 115 + 1 UUCCAAUGCGAU-----UUACUUACCAAAUUGCAUAUUCGUGAGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCA .....(((((((-----((.......)))))))))....(((.(((((.......((....)).....)))))...)))....((((..((((((((.((....)))))))))).)))). ( -45.30) >DroSim_CAF1 9516 115 + 1 UUCUAAUGCGAU-----UUACUUACCAAAUUGCAUAUUCGUGUGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCA .....(((((((-----((.......)))))))))....(((((((((.......((....)).....))))))..)))....((((..((((((((.((....)))))))))).)))). ( -46.80) >DroEre_CAF1 8239 120 + 1 UCCUUAUGCGAUUAAAUUUGCUUACCAAGUUGCAUAUUCGUGUGCCAGAUUUCUUCGGCGACGUCUCCCCGGCAGCCACAGCAGCCGCAGAGGAGGCAGGAUCGCCGCUGCUUCCUGGCA ....(((((((((..............)))))))))......((((((....(..((((((.(((((((((((.((....)).))))..).))))))....))))))..)....)))))) ( -41.84) >consensus UUCUAAUGCGAU_____UUACUUACCAAAUUGCAUAUUCGUGUGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCA .....(((((((................)))))))........(((((.......((....))......((((.((....)).))))..((((((((.((....))))))))))))))). (-36.73 = -35.66 + -1.06)

| Location | 11,380,867 – 11,380,987 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.53 |
| Mean single sequence MFE | -53.55 |
| Consensus MFE | -46.49 |
| Energy contribution | -48.17 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
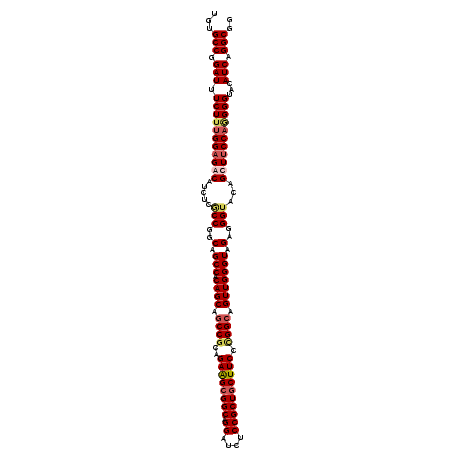
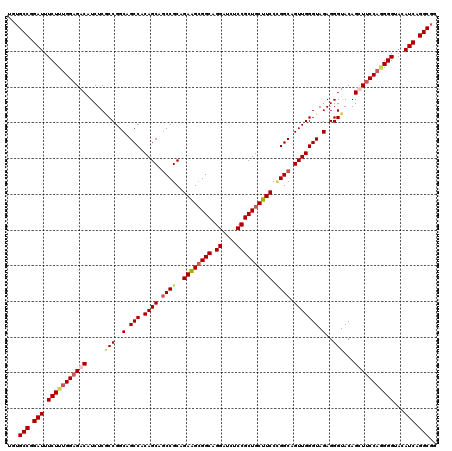
>2R_DroMel_CAF1 11380867 120 + 20766785 CGUGCCUGAUUUCUUUGGAGACAUCUCACCGGCAGCCACAGCCCCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCAGGCAGUUGGGUAGAGGGUACAGCUUCCAAGGGUACAUCAGGCGG ..((((((((.(((((((((((.((((.(((((.(((...((....)).((((((((.((....))))))))))..))).)))))...)))))).....)))))))))...)))))))). ( -52.60) >DroSec_CAF1 8128 120 + 1 UGAGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCAGUUGGGUAGAGGGUACAGUUUCCAGGGGUAUAUCAGGCGG ...(((.(((.(((((((((((.....(((..(.(((.((((.((((..((((((((.((....)))))))))).)))).))))))).)..)))...)))))))))))...))).))).. ( -56.90) >DroSim_CAF1 9551 120 + 1 UGUGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCAGUUGGGUAGAGGGUACAGUUUCCAGGGGUAUAUCAGGCGG ..((((.(((.(((((((((((.....(((..(.(((.((((.((((..((((((((.((....)))))))))).)))).))))))).)..)))...)))))))))))...))).)))). ( -56.90) >DroEre_CAF1 8279 120 + 1 UGUGCCAGAUUUCUUCGGCGACGUCUCCCCGGCAGCCACAGCAGCCGCAGAGGAGGCAGGAUCGCCGCUGCUUCCUGGCAGUUGGGUAGAGGGUACAGCUUCCAUGGGCACAUCAGGCGG ((((((.((.....))(((..(.(((.((((((.(((...((....))..(((((((((........)))))))))))).)))))).))).).....)))......))))))........ ( -47.80) >consensus UGUGCCGGAUUUCUUUGGAGACAUCUCGCCGGCAGCCACAGCAGCCGCAGAAGCGGCAGGAUCUCCGCUGCUUCCCGGCAGUUGGGUAGAGGGUACAGCUUCCAGGGGUACAUCAGGCGG ...(((.(((.(((((((((((.....(((..(.(((.((((.((((..((((((((.((....)))))))))).)))).))))))).)..)))...)))))))))))...))).))).. (-46.49 = -48.17 + 1.69)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:32 2006