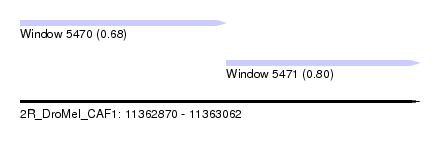
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,362,870 – 11,363,062 |
| Length | 192 |
| Max. P | 0.795572 |

| Location | 11,362,870 – 11,362,969 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -32.40 |
| Consensus MFE | -10.92 |
| Energy contribution | -10.32 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.34 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.680453 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11362870 99 + 20766785 AGCAACAAAUAUUUGC------UAGUGGCAUCCAAAAUGUGUUUCGAAAAACCGUUGA------CUGUAGCAAUUACCUGCUGUCAGCUAGCUUGGAGCGUCGCAGCAGUG ............((((------(.(((((.(((((...(.((((....)))))(((((------(.((((.......)))).))))))....)))))..)))))))))).. ( -28.90) >DroSec_CAF1 21032 97 + 1 AGCAACAAAUAUUUGC------UAGUGGCAUCUAAA--GUGUUUCAAAAAACCGUUGG------CUGUAGCAAUUACCAGCUGUCAGCUAGCUCGGAGCAUCACAGCAGUG ............((((------(.((((..(((...--(.((((....)))))(((((------(((((((........)))).))))))))..)))...))))))))).. ( -26.80) >DroSim_CAF1 20777 97 + 1 AGCAACAAAUAUUUGC------AAGUGGCAUCUAAA--GUGUUUCAAAAAACCGUUGG------CUGUAGCAACUACCUGCUGUCAGCUAGCUCGGAGCGUCACAGCAGUG .((((.......))))------..(((((.(((...--(.((((....)))))(((((------((((((((......))))).))))))))..)))..)))))....... ( -29.50) >DroEre_CAF1 23258 103 + 1 AGCAACAAAUAUUCGC------UGCUGACAGCUAAA--GCAAUUCAAAAAACCGUUGAAGCGGACUGUAGCAGGUACCUGCUGUCAGCUUGCGCGGACUGUCGCAGCAGUG ..............((------(((.(((((((...--((((.........((((....)))).(((((((((....)))))).))).))))..)).)))))))))).... ( -38.80) >DroYak_CAF1 21446 109 + 1 AGCAACAAAUAUUUGCUUUAGCUAUUGACAUCUAAA--GUAAUUUGAAAAACCGUUGAAGCGCUCUGUAGCAGUUAGCUGGUGUCAGCUUGCUCGGAGCGUCGCAGCAGUG .....(((((...((((((((..........)))))--))))))))....((.((((..((((((((.((((...(((((....)))))))))))))))))..)))).)). ( -38.00) >consensus AGCAACAAAUAUUUGC______UAGUGGCAUCUAAA__GUGUUUCAAAAAACCGUUGA______CUGUAGCAAUUACCUGCUGUCAGCUAGCUCGGAGCGUCGCAGCAGUG .((.(((..................((.....)).........((((.......)))).......))).))......(((((((..(((.......)))...))))))).. (-10.92 = -10.32 + -0.60)

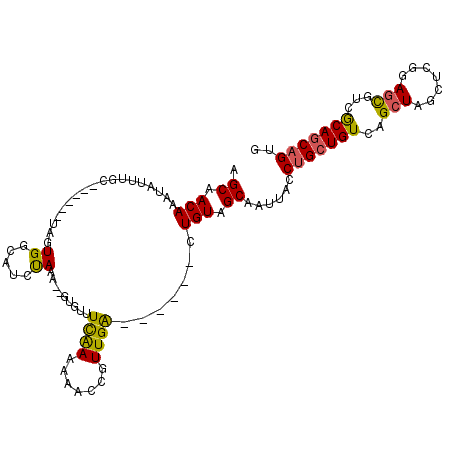
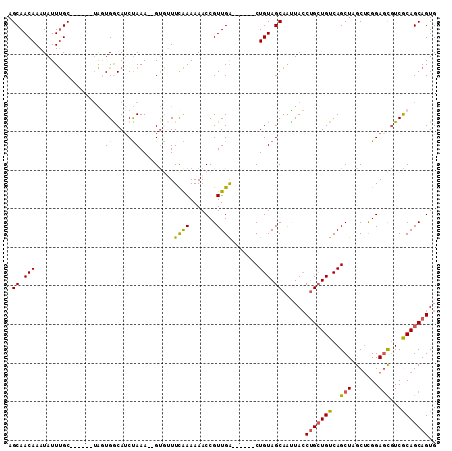
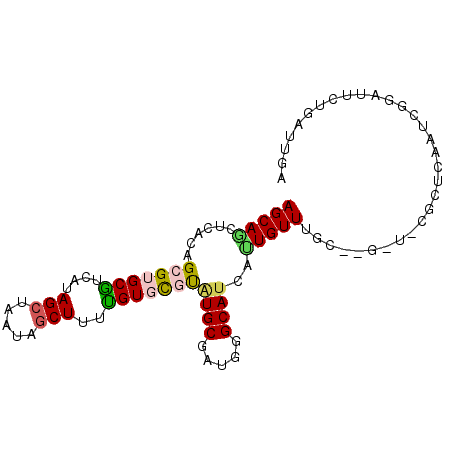
| Location | 11,362,969 – 11,363,062 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 74.54 |
| Mean single sequence MFE | -27.04 |
| Consensus MFE | -10.18 |
| Energy contribution | -10.30 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.37 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.38 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.795572 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11362969 93 + 20766785 AGCAGCUCACAGCGUGCAACAUAGCUAAUAGCUUUUGUGCGUAUGCGAUGGGCAUCACUGUUUGCUUGCUGCGCUCAAUUGGAUUCUGAUUGA .(((((.....((((((.(((.(((.....)))..)))..))))))...(((((........))))))))))..(((((..(...)..))))) ( -30.80) >DroSec_CAF1 21129 84 + 1 AGCAGCUCACAGCGUGCGUCAUAGCUAAAAGCUUUUGUGCGUAUGCGAUGGGCAUCAUUGUU---------CGCUCAAUCGGAUUCUGAUUGA ....(((((..((((((((((.(((.....)))..)).))))))))..))))).........---------...((((((((...)))))))) ( -29.80) >DroSim_CAF1 20874 84 + 1 AGCAGCUCACAGCGUGCGUCAUAGCUAAAAGCUUUUGUGCGUGUGCGAUGGGCAUCAUUGUU---------CGCUCAAUCGGAUUCUGAUUGA ....(((((..(((..(((((.(((.....)))..)).)))..)))..))))).........---------...((((((((...)))))))) ( -28.40) >DroEre_CAF1 23361 78 + 1 AGCAAUCCAC-GCGUGCUUCGUAGCUAAUAGCUUUAGUCCUCGUGCGAUGGGCAUCAUUGUUUGCCCGUU--------------UCUGAUUGG ..((((((((-(.(..((....(((.....)))..))..).)))).((((((((........))))))))--------------...))))). ( -25.80) >DroYak_CAF1 21555 93 + 1 AGCAAUGCACAGCGCGCUUUAUAGCUUAUAACUUUAGUCUCUAUGCGAUGGGCAGCACUGUUUGCUUGCUUCGCUAAAUAGAAUUCUGAUUGG (((((.((((((.((((((....((.....((....))......))...)))).)).))))..)))))))...((((.(((....))).)))) ( -20.40) >consensus AGCAGCUCACAGCGUGCGUCAUAGCUAAUAGCUUUUGUGCGUAUGCGAUGGGCAUCAUUGUUUGC__G_U_CGCUCAAUCGGAUUCUGAUUGA (((((......(((((((....(((.....)))..)))))))((((.....))))..)))))............................... (-10.18 = -10.30 + 0.12)

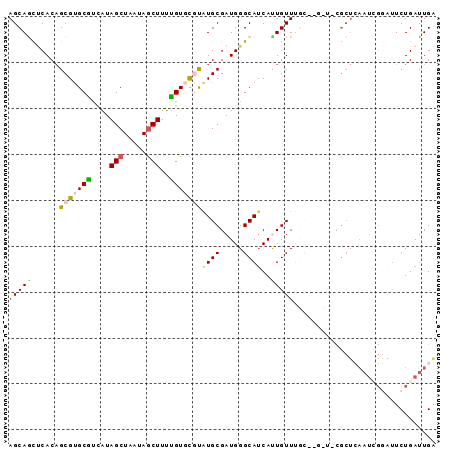
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:16 2006