
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,349,473 – 11,349,565 |
| Length | 92 |
| Max. P | 0.823299 |

| Location | 11,349,473 – 11,349,565 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 86.99 |
| Mean single sequence MFE | -19.27 |
| Consensus MFE | -13.71 |
| Energy contribution | -13.78 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.823299 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
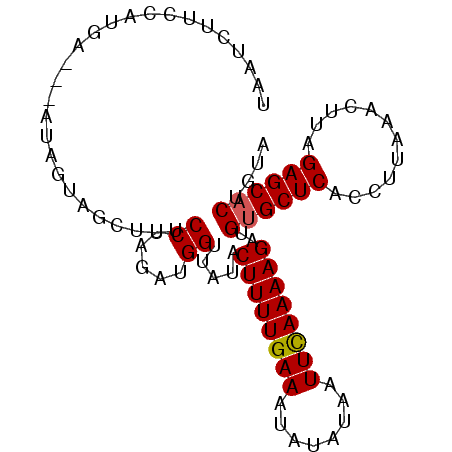
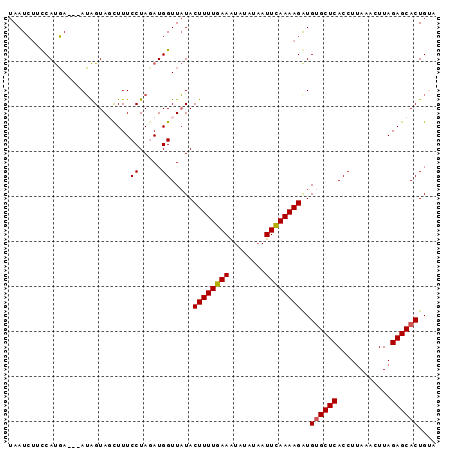
>2R_DroMel_CAF1 11349473 92 + 20766785 UAAUCUUCCAUGA---AUAGUAUCUUUCCUAGAUGGUUAUACUUUUGAAAUAUAUAAUUCAAAAGAUGUGCUCACCUUAAACUUAGAGCACUGAA (((((.((...((---(........)))...)).)))))..((((((((........))))))))..((((((............)))))).... ( -18.00) >DroSec_CAF1 7655 91 + 1 UAAUCUUCCAUGA---AUAGUAGCUUUCCUAUAUGGUUAUACUUUUGAAAUAUAUAAUUCAAAAGGUGUGCUCACCUUAAACUUAGAGCACUG-A .......((((..---((((........)))))))).....((((((((........))))))))..((((((............))))))..-. ( -18.20) >DroSim_CAF1 7479 91 + 1 UAAUCUUCCAUGA---AUAGUAGCUUUCCUAGAUGGUUAUACUUUUGAAAUAUAUAAUUCAAAAGGUG-GCUCACCUUAAACUUAGAGCACUGUA .............---(((((.(((((...((.(((((((.((((((((........)))))))))))-)).)).)).......)))))))))). ( -18.30) >DroYak_CAF1 7545 95 + 1 UAAUAACUCACUAAAGAGGGUAUUUUUCCUAGAUGGUUCUACUUUUGAAGUACAUCAUUUAAAAGAUGUGCUCACCUCAAACUUAGAGCACUGUA ................((((......)))).....((((((..(((((((((((((........))))))))....)))))..))))))...... ( -22.60) >consensus UAAUCUUCCAUGA___AUAGUAGCUUUCCUAGAUGGUUAUACUUUUGAAAUAUAUAAUUCAAAAGAUGUGCUCACCUUAAACUUAGAGCACUGUA ...........................((.....)).....((((((((........))))))))..((((((............)))))).... (-13.71 = -13.78 + 0.06)

| Location | 11,349,473 – 11,349,565 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 86.99 |
| Mean single sequence MFE | -19.15 |
| Consensus MFE | -13.80 |
| Energy contribution | -14.30 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.669951 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
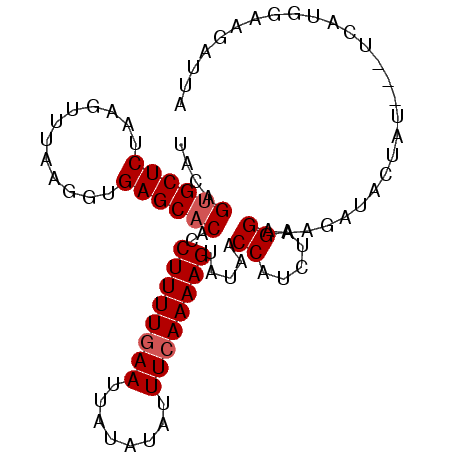
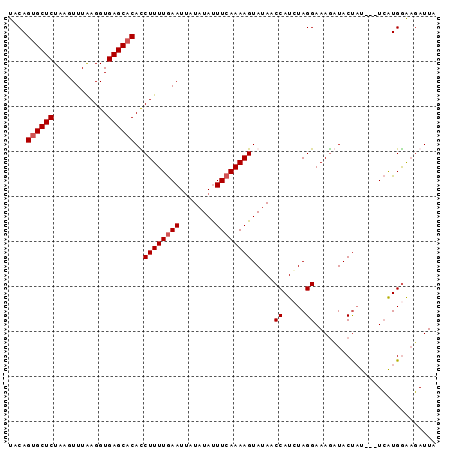
>2R_DroMel_CAF1 11349473 92 - 20766785 UUCAGUGCUCUAAGUUUAAGGUGAGCACAUCUUUUGAAUUAUAUAUUUCAAAAGUAUAACCAUCUAGGAAAGAUACUAU---UCAUGGAAGAUUA ....((((((............))))))..((((((((........)))))))).....((((.(((........))).---..))))....... ( -20.00) >DroSec_CAF1 7655 91 - 1 U-CAGUGCUCUAAGUUUAAGGUGAGCACACCUUUUGAAUUAUAUAUUUCAAAAGUAUAACCAUAUAGGAAAGCUACUAU---UCAUGGAAGAUUA .-..((((((............))))))..((((((((........)))))))).....((((((((........))))---..))))....... ( -21.60) >DroSim_CAF1 7479 91 - 1 UACAGUGCUCUAAGUUUAAGGUGAGC-CACCUUUUGAAUUAUAUAUUUCAAAAGUAUAACCAUCUAGGAAAGCUACUAU---UCAUGGAAGAUUA ....((((((............))).-)))((((((((........)))))))).....((((.(((........))).---..))))....... ( -15.50) >DroYak_CAF1 7545 95 - 1 UACAGUGCUCUAAGUUUGAGGUGAGCACAUCUUUUAAAUGAUGUACUUCAAAAGUAGAACCAUCUAGGAAAAAUACCCUCUUUAGUGAGUUAUUA ........((((..((((((((....(((((........)))))))))))))..)))).((.....)).........(((......)))...... ( -19.50) >consensus UACAGUGCUCUAAGUUUAAGGUGAGCACACCUUUUGAAUUAUAUAUUUCAAAAGUAUAACCAUCUAGGAAAGAUACUAU___UCAUGGAAGAUUA ....((((((............))))))..((((((((........)))))))).....((.....))........................... (-13.80 = -14.30 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:08 2006