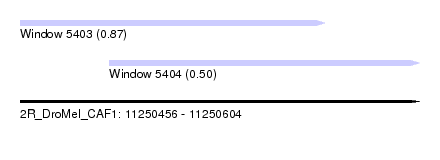

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,250,456 – 11,250,604 |
| Length | 148 |
| Max. P | 0.870300 |

| Location | 11,250,456 – 11,250,569 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.18 |
| Mean single sequence MFE | -40.15 |
| Consensus MFE | -26.33 |
| Energy contribution | -25.97 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870300 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11250456 113 + 20766785 ACUGUACGACGAUUCG------AGUAGCAGCAG-CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUCUAUGCCGAAUUUCGCGCGAGUGAAAGCAGAUUCGAACGGGGUC .((((..((((.((((------(((.(((((..-...))))).(((.(((....))).)))...))))))))))).......((((((((((....)))).....)))))).)))).... ( -44.20) >DroVir_CAF1 4045 114 + 1 --CGUGU---AGCAGCAGUAGCAGCAGCAGCGA-CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAACGCUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUCUCUUCGAGCAUCGCC --.....---.((((((((.((.......)).)-).))))))(((...((((.(((((((......))))))).))))((((((((((((((....)))))).....)))).)))).))) ( -38.20) >DroPse_CAF1 40359 111 + 1 --CGUAU---AAAAACAGC---AGCGACGGCAG-CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUGUACGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGGGGAA --.....---.......((---(((.(((....-))))))))..(((.(((.((((((((......)))))))).))).(((((((((((((....)))).....)))))).)))))).. ( -38.10) >DroGri_CAF1 4022 114 + 1 --CGUGU---CGCAGCAGCAACAGCAGCAGCGA-CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUCUCUUCGAGCAUCGCC --...((---(((.((.((....)).)).))))-)(((.((((((...((((((((((((......))))))))))))..)))((.((((((....)))))).)).......))).))). ( -45.40) >DroWil_CAF1 68031 103 + 1 -UCGUUU---GCGU----------CGUCGUCGGUGGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGUG--- -.(((((---(.((----------(....(((((((.((....)))).((((((((((((......))))))))))))..))))).((((((....))))))...))).))))))..--- ( -36.90) >DroPer_CAF1 39872 111 + 1 --CGUAU---AAAAACAGC---AGCGACGGCAG-CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUGUACGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGGGGAA --.....---.......((---(((.(((....-))))))))..(((.(((.((((((((......)))))))).))).(((((((((((((....)))).....)))))).)))))).. ( -38.10) >consensus __CGUAU___AAAAACAGC___AGCAGCAGCAG_CGUGCUGCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUCUGUAUGCCGAAUUUCGCGCGAGUGAAAGUAGAUUCGAGCGGGGAC .......................((..(((((....)))))((((...(((.((((((((......)))))))).)))..))))..((((((....))))))..........))...... (-26.33 = -25.97 + -0.36)


| Location | 11,250,489 – 11,250,604 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.37 |
| Mean single sequence MFE | -35.35 |
| Consensus MFE | -21.67 |
| Energy contribution | -21.75 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.61 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 11250489 115 + 20766785 GCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUCUAUGCCGAAUUUCGCGCGAGUGAAAGCAGAUUCGAACGGGGUCUGAUCCAGU-----GCCAGUUGGACAUUGAAAACGCCAAC ..(((......((((((..((.....(((.((((....))))))).((((((....)))))).(((((((.....))))))).......-----....))..))))))......)))... ( -36.70) >DroVir_CAF1 4079 115 + 1 GCGGCCCAACAAAUGUUUGGCAGAACUCGAACGCUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUCUCUUCGAGCAUCGCCUGAGCCAGG----CGCCAA-ACGAUAACGAUAUCGAUAAC ..............(((((((.((.((((((.((......)).((.((((((....)))))).))..))))))..))(((((...))))----))))))-))((((....))))...... ( -38.20) >DroPse_CAF1 40390 112 + 1 GCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUGUACGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGGGGAACGGG---GC-----GCCAGUCGGAUAUUGAUAACGCCAAC ..(((......((((((((((.(....)(.((((((...(((((((((((((....)))).....)))))).)))(....).))---))-----))).))))))))))......)))... ( -36.60) >DroGri_CAF1 4056 118 + 1 GCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUCUCUUCGAGCAUCGCCUCAGUCAGAGUAACGAC-A-ACGAUACCGAUAACGAUAAC .((((...((((((((((((......))))))))))))..))))..((((((....))))))(((...(((.(.((((.....(((........)))-.-.)))).))))....)))... ( -33.70) >DroWil_CAF1 68057 92 + 1 GCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUUUGUAUGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGUG-----AACUAGU-----G--A----------------GUCAAC .((((...((((((((((((......))))))))))))..))))..((((((....))))))...((((((...(..-----..)...)-----)--)----------------)))... ( -29.70) >DroPer_CAF1 39903 112 + 1 GCGGCCCAACAAAUGUUUGGCAGAACUCGAGCGUCUGUACGCCGAAUUUCGCGCGAGUGAAAGUAGAUUUGAGCGGGGAACGGG---AC-----GCCAGUCGGAUAUUGAUAACGCCAAC ..(((......((((((((((.(....)(.((((((...(((((((((((((....)))).....)))))).)))(....).))---))-----))).))))))))))......)))... ( -37.20) >consensus GCGGCCCAACAAAUGUUUGGCAGAACUCGAACGUCUGUAUGCCGAAUUUCGCGCGAGUGAAAGUAGAUUCGAGCGGGGACUGAGCCAGC_____GCCAG_AGGAUAUUGAUAACGCCAAC .((((...(((.((((((((......)))))))).)))..))))..((((((....)))))).......................................................... (-21.67 = -21.75 + 0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:11 2006