
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 11,224,433 – 11,224,539 |
| Length | 106 |
| Max. P | 0.652531 |

| Location | 11,224,433 – 11,224,539 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.22 |
| Mean single sequence MFE | -23.68 |
| Consensus MFE | -18.91 |
| Energy contribution | -18.93 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652531 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
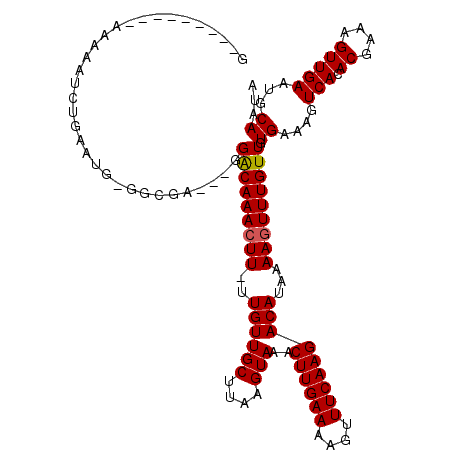
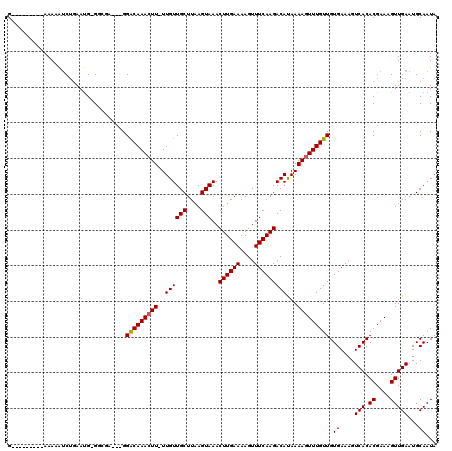
>2R_DroMel_CAF1 11224433 106 - 20766785 G---------AAAAAUCUGAAUG-GGCGA---GGACAAACUU-UUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACAUAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .---------.............-.((((---.(((((((((-(((((((....)))..((((((....))))))..))))))))))))).).....(((.((....))))).))).... ( -24.30) >DroPse_CAF1 13286 105 - 1 G---------AAAAAUCUGAAUG-GU--A---GGACAAACUUUUUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACACAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .---------.......(((...-.(--(---.(((((((((((((((((....)))..((((((....))))))))).))))))))))).))....))).((....))........... ( -22.20) >DroWil_CAF1 28408 116 - 1 GUCGAAAAACAAAAGGCGAAACGUGGAAA---GGGCAAACUU-UUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACAUAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA ...............(((....((((..(---.(((((((((-(((((((....)))..((((((....))))))..))))))))))))).).....))))((....))....))).... ( -25.90) >DroYak_CAF1 11797 106 - 1 G---------AAAAAUCUGAAUG-GGCGA---GGACAAAGUU-UUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACAUAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .---------.............-.((((---.((((((.((-(((((((....)))..((((((....))))))..)))))).)))))).).....(((.((....))))).))).... ( -20.40) >DroAna_CAF1 11851 109 - 1 G---------AAAAAUCUGAAUG-GGCGAGCAGGACAAACUU-UUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACAUAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .---------.............-.(((..((.(((((((((-(((((((....)))..((((((....))))))..))))))))))))).))....(((.((....))))).))).... ( -27.10) >DroPer_CAF1 12844 105 - 1 G---------AAAAAUCUGAAUG-GU--A---GGACAAACUUUUUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACACAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .---------.......(((...-.(--(---.(((((((((((((((((....)))..((((((....))))))))).))))))))))).))....))).((....))........... ( -22.20) >consensus G_________AAAAAUCUGAAUG_GGCGA___GGACAAACUU_UUGUUGCUUAAGUAAACUUGAAAAGUUUCAAGACAUAAAAGUUUGUUGUGAAAGUCACACGAAAGUUGAAUGCAAUA .................................(((((((((..((((((....)))..((((((....)))))))))...))))))))).((....(((.((....)))))...))... (-18.91 = -18.93 + 0.03)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:52 2006