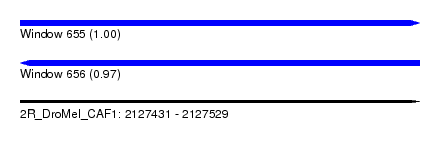
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,127,431 – 2,127,529 |
| Length | 98 |
| Max. P | 0.996726 |

| Location | 2,127,431 – 2,127,529 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 94.69 |
| Mean single sequence MFE | -30.47 |
| Consensus MFE | -27.29 |
| Energy contribution | -27.01 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.74 |
| SVM RNA-class probability | 0.996726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
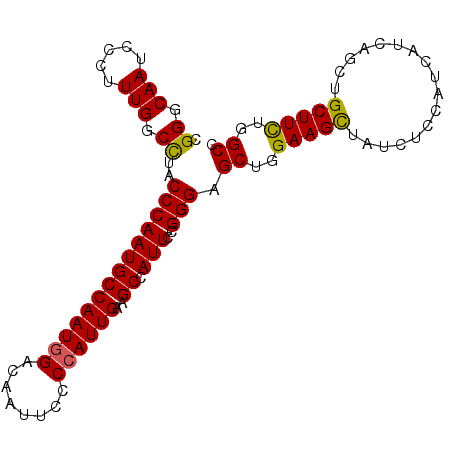
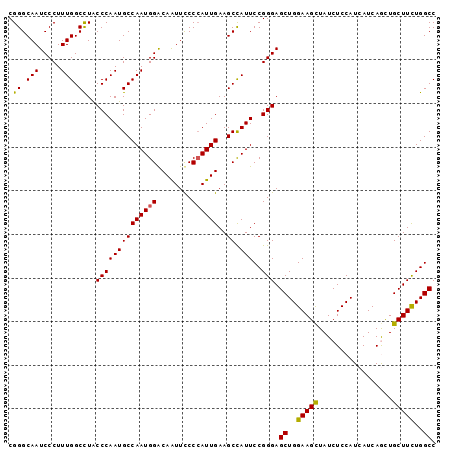
>2R_DroMel_CAF1 2127431 98 + 20766785 CGGGCAAUCCCUUUGGCUUACCCAAUGCCAAUGGGCAAUUCCCCAUUGAAGCCAUUCCGGGAGCUGGAAGCUAUCUCCAUCAUCAGCUGCUUUUGGCC .......((((..((((((.........(((((((......)))))))))))))....))))((..(((((...((........))..)))))..)). ( -31.80) >DroSec_CAF1 106102 98 + 1 CGGGCAAUCCCUUUGGCCUGCCCAAUGCCAAUGGGCAAUUCCCCAUUGAAGCCAUUCCGGGAGCUGGAAGCUAUCUCCAUCAUCAGCUGCUUCUGGCC .(((((..((....))..)))))...(((((((((......))))))(((((...(((((...)))))((((............))))))))).))). ( -33.80) >DroSim_CAF1 110055 98 + 1 CGGGCAAUCCCUUUGGCCUACCCAAUGCCAAUGGACAAUUCCCCAUUGAAGCCAUUCCGGGAGCUGGAAGCUAUCUCCAUCAUCAGCUGCUUCUGGCC .((.(((.....))).))..((((((((((((((........))))))..).))))..))).((..(((((...((........))..)))))..)). ( -29.00) >DroEre_CAF1 104830 98 + 1 CGGGCAAUCCCUUUGGCCUACCCAAUGCCAAUGGACAGUUUCCAAUUGAAGCUAUUCCGGGAGCUGGAAGCUAUCUCCAUCAUCAACUGCUUCUGGCC .......((((.(((((.........))))).(((.((((((.....))))))..)))))))((..(((((.................)))))..)). ( -30.63) >DroYak_CAF1 107132 98 + 1 CGGGCAAUCCCUUUGGCCUACCCAAUGCCAAUGGACAGUUUCCCAUUGAAGCCAUUCCGGGAGCUGGAAGUUAUCUCCAUCAUCAAUUGCUUCUGGCC .......((((.(((((.........))))).(((..(((((.....)))))...)))))))((..(((((.................)))))..)). ( -27.13) >consensus CGGGCAAUCCCUUUGGCCUACCCAAUGCCAAUGGACAAUUCCCCAUUGAAGCCAUUCCGGGAGCUGGAAGCUAUCUCCAUCAUCAGCUGCUUCUGGCC .((.(((.....))).))..((((((((((((((........))))))..)).)))..))).((..(((((.................)))))..)). (-27.29 = -27.01 + -0.28)

| Location | 2,127,431 – 2,127,529 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 94.69 |
| Mean single sequence MFE | -33.92 |
| Consensus MFE | -33.02 |
| Energy contribution | -32.62 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.972501 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
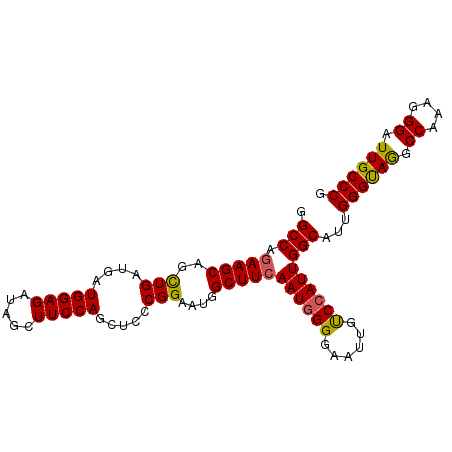
>2R_DroMel_CAF1 2127431 98 - 20766785 GGCCAAAAGCAGCUGAUGAUGGAGAUAGCUUCCAGCUCCCGGAAUGGCUUCAAUGGGGAAUUGCCCAUUGGCAUUGGGUAAGCCAAAGGGAUUGCCCG .(((..((((..(((....(((((.....))))).....)))....)))).((((((......)))))))))...((((((.((....)).)))))). ( -33.20) >DroSec_CAF1 106102 98 - 1 GGCCAGAAGCAGCUGAUGAUGGAGAUAGCUUCCAGCUCCCGGAAUGGCUUCAAUGGGGAAUUGCCCAUUGGCAUUGGGCAGGCCAAAGGGAUUGCCCG .(((.(((((..(((....(((((.....))))).....)))....)))))((((((......)))))))))...((((((.((....)).)))))). ( -39.30) >DroSim_CAF1 110055 98 - 1 GGCCAGAAGCAGCUGAUGAUGGAGAUAGCUUCCAGCUCCCGGAAUGGCUUCAAUGGGGAAUUGUCCAUUGGCAUUGGGUAGGCCAAAGGGAUUGCCCG .(((.(((((..(((....(((((.....))))).....)))....)))))((((((......)))))))))...((((((.((....)).)))))). ( -34.40) >DroEre_CAF1 104830 98 - 1 GGCCAGAAGCAGUUGAUGAUGGAGAUAGCUUCCAGCUCCCGGAAUAGCUUCAAUUGGAAACUGUCCAUUGGCAUUGGGUAGGCCAAAGGGAUUGCCCG .(((((..((((((..((((((((.....((((.......))))...)))).))))..))))))...)))))...((((((.((....)).)))))). ( -31.70) >DroYak_CAF1 107132 98 - 1 GGCCAGAAGCAAUUGAUGAUGGAGAUAACUUCCAGCUCCCGGAAUGGCUUCAAUGGGAAACUGUCCAUUGGCAUUGGGUAGGCCAAAGGGAUUGCCCG .(((.(((((..(((....(((((.....))))).....)))....)))))((((((......)))))))))...((((((.((....)).)))))). ( -31.00) >consensus GGCCAGAAGCAGCUGAUGAUGGAGAUAGCUUCCAGCUCCCGGAAUGGCUUCAAUGGGGAAUUGUCCAUUGGCAUUGGGUAGGCCAAAGGGAUUGCCCG .(((.(((((..(((....(((((.....))))).....)))....)))))((((((......)))))))))...((((((.((....)).)))))). (-33.02 = -32.62 + -0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:35:29 2006