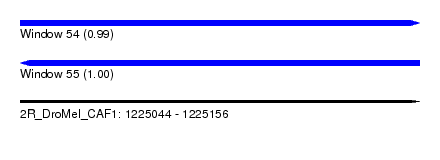

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 1,225,044 – 1,225,156 |
| Length | 112 |
| Max. P | 0.999965 |

| Location | 1,225,044 – 1,225,156 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 97.56 |
| Mean single sequence MFE | -35.90 |
| Consensus MFE | -34.60 |
| Energy contribution | -34.80 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.29 |
| SVM RNA-class probability | 0.991815 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
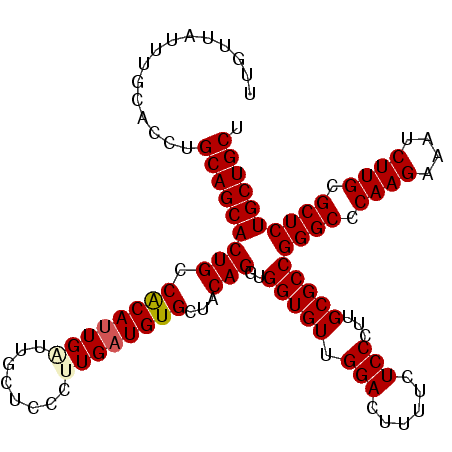
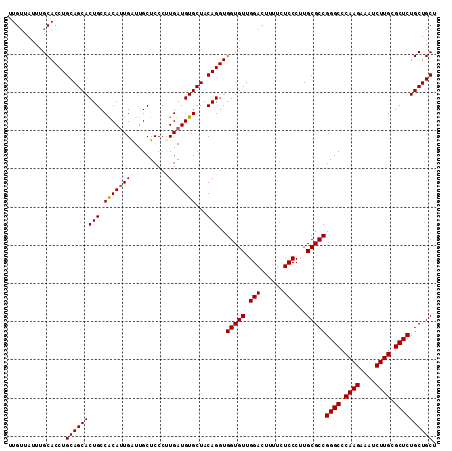
>2R_DroMel_CAF1 1225044 112 + 20766785 UUGUUAUUUGCACCUGCAGCACUGCCACAUUGAUUGCUCCCUUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.(((((..(........)..)))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). ( -36.60) >DroVir_CAF1 28338 112 + 1 UUGUUAUUUGCACCUGCAGCACUGCCGCAUUGAUUGCUCCCUUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.(((((..(........)..)))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). ( -36.20) >DroPse_CAF1 17250 112 + 1 UUGUUAUUUGCACCUGCAGCACUGCCACAUUGAUUGCUCCCCUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.((((((............))))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). ( -34.30) >DroGri_CAF1 19182 112 + 1 UUGUUAUUUGCACCUGCAGCACUGCCGCAUUGAUUGCUCCCUUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.(((((..(........)..)))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). ( -36.20) >DroAna_CAF1 17014 112 + 1 UUGGUACUUGCACCUGCAGCACUGCCACAGUGGUUGCUCCCUUGAUGUGCUACAGGUGGUGUUGGACUUUUUUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ..((((...(((((((.(((((...((.((.((.....))))))..))))).)))))(((((.(((......)))...)))))((((.((((....)))).)))).)))))) ( -37.80) >DroPer_CAF1 17285 112 + 1 UUGUUAUUUGCACCUGCAGCACUGCCACAUUGAUUGCUCCCCUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.((((((............))))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). ( -34.30) >consensus UUGUUAUUUGCACCUGCAGCACUGCCACAUUGAUUGCUCCCUUGAUGUGCUACAGGUGGUGUUGGACUUUUCUCCCUUGCGCCGGGCCCAAGAAAUCUUGCGCUCUGCUGCU ...............(((((((((.((((((((........))))))))...)))..(((((.(((......)))...)))))((((.((((....)))).)))))))))). (-34.60 = -34.80 + 0.20)

| Location | 1,225,044 – 1,225,156 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 97.56 |
| Mean single sequence MFE | -39.57 |
| Consensus MFE | -38.24 |
| Energy contribution | -38.02 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.97 |
| SVM decision value | 4.96 |
| SVM RNA-class probability | 0.999965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
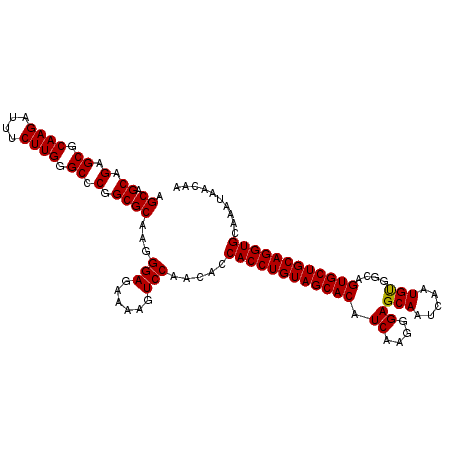
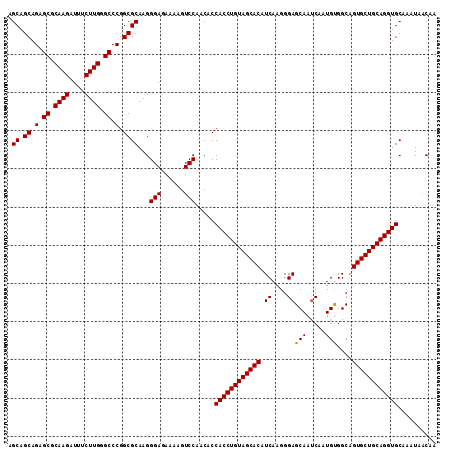
>2R_DroMel_CAF1 1225044 112 - 20766785 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAAGGGAGCAAUCAAUGUGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.(((...((....))....)))..)))))))))))).......... ( -40.20) >DroVir_CAF1 28338 112 - 1 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAAGGGAGCAAUCAAUGCGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.((....))(((.....)))....)))))))))))).......... ( -39.50) >DroPse_CAF1 17250 112 - 1 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAGGGGAGCAAUCAAUGUGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.(((.(.((....))..).)))..)))))))))))).......... ( -39.30) >DroGri_CAF1 19182 112 - 1 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAAGGGAGCAAUCAAUGCGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.((....))(((.....)))....)))))))))))).......... ( -39.50) >DroAna_CAF1 17014 112 - 1 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAAAAAAGUCCAACACCACCUGUAGCACAUCAAGGGAGCAACCACUGUGGCAGUGCUGCAGGUGCAAGUACCAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.(((((((.....)).)).)))..)))))))))))).......... ( -39.60) >DroPer_CAF1 17285 112 - 1 AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAGGGGAGCAAUCAAUGUGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.(((.(.((....))..).)))..)))))))))))).......... ( -39.30) >consensus AGCAGCAGAGCGCAAGAUUUCUUGGGCCCGGCGCAAGGGAGAAAAGUCCAACACCACCUGUAGCACAUCAAGGGAGCAAUCAAUGUGGCAGUGCUGCAGGUGCAAAUAACAA .((.((.(.((.((((....)))).)).).))))...(((......))).....((((((((((((.((....))(((.....)))....)))))))))))).......... (-38.24 = -38.02 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:23:58 2006