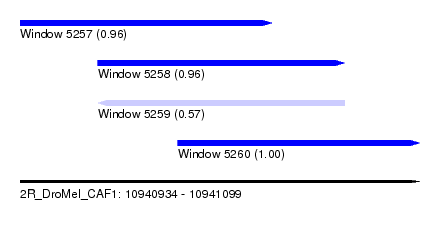

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,940,934 – 10,941,099 |
| Length | 165 |
| Max. P | 0.998532 |

| Location | 10,940,934 – 10,941,038 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 99.23 |
| Mean single sequence MFE | -32.40 |
| Consensus MFE | -32.40 |
| Energy contribution | -32.40 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.63 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10940934 104 + 20766785 AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) ( -32.40) >DroSec_CAF1 53649 104 + 1 AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCUGAAUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) ( -32.40) >DroSim_CAF1 55663 104 + 1 AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) ( -32.40) >DroEre_CAF1 55206 104 + 1 AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGACUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) ( -32.40) >DroYak_CAF1 55803 104 + 1 AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) ( -32.40) >consensus AUGUAAAAAUCUAUCUAGAGCUAGAGGCUCGGCUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAA .................((((.....))))((.((((.((((.....))))....)))))).(((((.(((((((.(((.........)))))))))).))))) (-32.40 = -32.40 + -0.00)

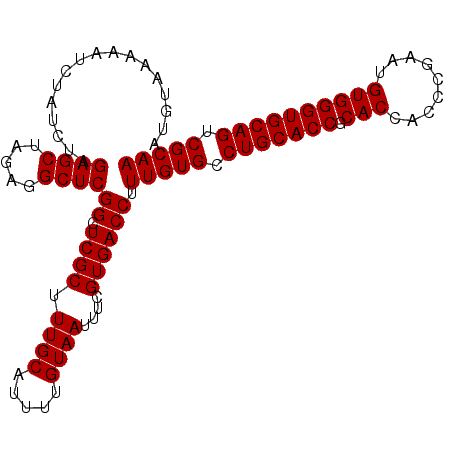
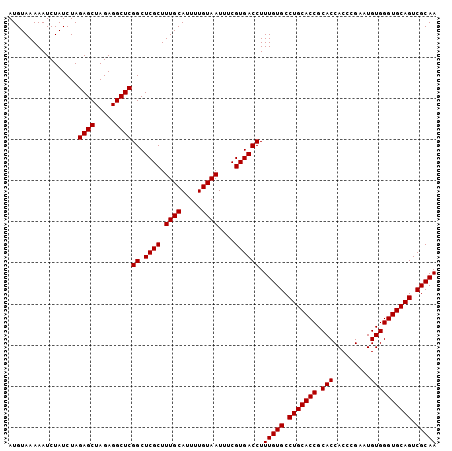
| Location | 10,940,966 – 10,941,068 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 98.82 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -27.80 |
| Energy contribution | -27.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.58 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10940966 102 + 20766785 CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACAA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... ( -27.80) >DroSec_CAF1 53681 102 + 1 CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCUGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... ( -27.80) >DroSim_CAF1 55695 102 + 1 CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... ( -27.80) >DroEre_CAF1 55238 102 + 1 CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGACUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... ( -27.80) >DroYak_CAF1 55835 102 + 1 CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... ( -27.80) >consensus CUCGCUUUGCAUUUUGUAAUUUCGUGACCUUUGUGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCA ......(((((...((((.....(((.(..(((((.(((((((.(((.........)))))))))).)))))..))))...))))......)))))...... (-27.80 = -27.80 + 0.00)
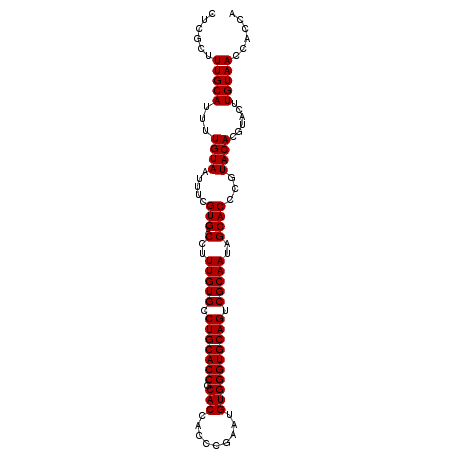


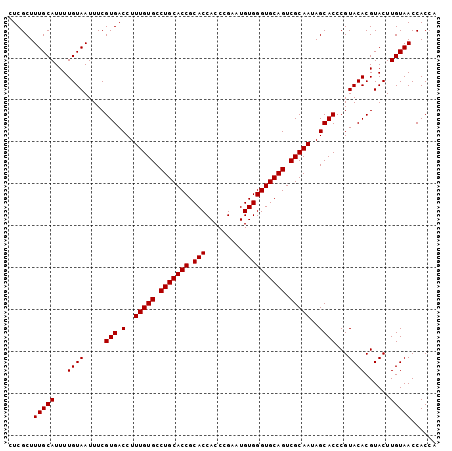
| Location | 10,940,966 – 10,941,068 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 98.82 |
| Mean single sequence MFE | -31.96 |
| Consensus MFE | -31.70 |
| Energy contribution | -31.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.573341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10940966 102 - 20766785 UUGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAUUCGGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ((((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...)))). ( -33.00) >DroSec_CAF1 53681 102 - 1 UGGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAUUCAGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ..((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...))... ( -31.70) >DroSim_CAF1 55695 102 - 1 UGGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAUUCGGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ..((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...))... ( -31.70) >DroEre_CAF1 55238 102 - 1 UGGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAGUCGGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ..((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...))... ( -31.70) >DroYak_CAF1 55835 102 - 1 UGGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAUUCGGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ..((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...))... ( -31.70) >consensus UGGUGGUUACAAGUACGUGUACGGGUGCUAUUGCGACUGCACCCACAUUCGGGUGGUGCGGUGCAGGCACAAAGGUCACGAAAUUACAAAAUGCAAAGCGAG ..((.((.....(((((((..(..(((((..(((.((((((((.((......))))))))))))))))))...)..))))....))).....))...))... (-31.70 = -31.70 + -0.00)
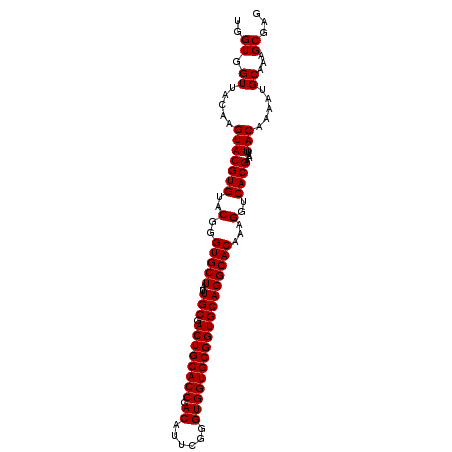


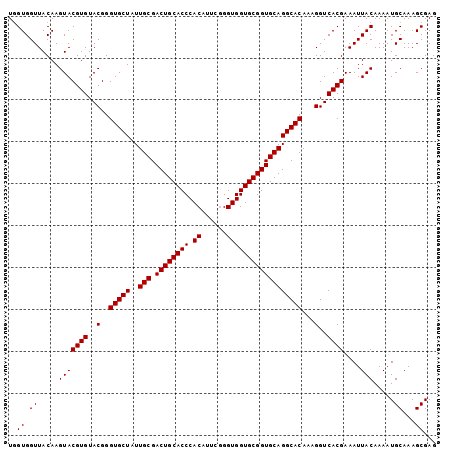
| Location | 10,940,999 – 10,941,099 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 92.73 |
| Mean single sequence MFE | -17.92 |
| Consensus MFE | -17.50 |
| Energy contribution | -17.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.98 |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.998532 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10940999 100 + 20766785 UGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACAAAAUAC------UACCAACUACCAACCACCCACA--ACAC ((((((((((.(((.........))))))))))..))).........(((....)))((((....)))).....------......................--.... ( -17.60) >DroSec_CAF1 53714 93 + 1 UGCCUGCACCGCACCACCUGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCAAAUAC------UAC-------CAACCACCCACA--ACAC ((((((((((.(((.........))))))))))..)))...............(((.(((.......))).)))------...-------............--.... ( -17.80) >DroSim_CAF1 55728 93 + 1 UGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCAAAUAC------UAC-------CAACCACCCACA--ACAC ((((((((((.(((.........))))))))))..)))...............(((.(((.......))).)))------...-------............--.... ( -17.80) >DroEre_CAF1 55271 93 + 1 UGCCUGCACCGCACCACCCGACUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCAAAUAC------UAC-------CAACCACCCACA--ACAC ((((((((((.(((.........))))))))))..)))...............(((.(((.......))).)))------...-------............--.... ( -17.80) >DroYak_CAF1 55868 101 + 1 UGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCAAAUACUACUACUAC-------CAACCACCCACAAAACAC ((((((((((.(((.........))))))))))..))).........(((...(((.(((.......))).)))...)))...-------.................. ( -18.60) >consensus UGCCUGCACCGCACCACCCGAAUGUGGGUGCAGUCGCAAUAGCACCCGUACACGUACUUGUAACCACCAAAUAC______UAC_______CAACCACCCACA__ACAC ((((((((((.(((.........))))))))))..)))...(((...(((....))).)))............................................... (-17.50 = -17.50 + 0.00)

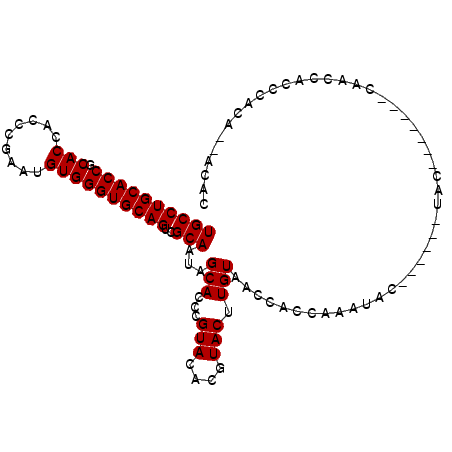

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:49 2006