
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,867,295 – 10,867,385 |
| Length | 90 |
| Max. P | 0.640120 |

| Location | 10,867,295 – 10,867,385 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 87.80 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -19.55 |
| Energy contribution | -22.05 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.610266 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10867295 90 + 20766785 AGCAUAUGUAGAGGCCACUCGGGCUCCUAUGUAUAUGCGU-UAUGUAGGAUCCAGGGUUCAGGGUCUUAACACUGGAUUAGAUGUCGAUCG .((((((((((..(((.....)))..))))..))))))..-.((((..(((((((.(((.((...)).))).)))))))..))))...... ( -27.30) >DroSec_CAF1 7825 87 + 1 AGCAUAUGUAGAGGCCAUUCGGGCUCCUA----UAUGCAUGUAUGUAGGAUCCAGGGUUCAGGGUCUUAACACUGGAUUAGAUGUCGAUCG .(((((((..(((.((....)).))).))----))))).((.((((..(((((((.(((.((...)).))).)))))))..)))))).... ( -26.50) >DroSim_CAF1 8181 91 + 1 AGCAUAUGUAGAGGCCAUUCGGGCUUCUAUGUAUAUGCAUGUAUGUAGGAUCCAGGGUUCAGGGUCUUAACACUGGAUUAGAUGUCGAUCG .(((((((((((((((.....)))))))))..)))))).((.((((..(((((((.(((.((...)).))).)))))))..)))))).... ( -31.60) >DroEre_CAF1 9147 77 + 1 AGCAUAUGCAGAGGCCAUUCGGGCUGCUA--------------UGUAGGAUCCAGGGUUCAGGGACUCAACACUGGAUUAGAUGUCGAUCG .(((((.((((...((....)).))))))--------------)))..(((((((.(((.((...)).))).))))))).(((....))). ( -21.80) >consensus AGCAUAUGUAGAGGCCAUUCGGGCUCCUA____UAUGCAU_UAUGUAGGAUCCAGGGUUCAGGGUCUUAACACUGGAUUAGAUGUCGAUCG .((((((((((.((((.....)))).))))..))))))......((..(((((((.(((.((...)).))).)))))))..))........ (-19.55 = -22.05 + 2.50)


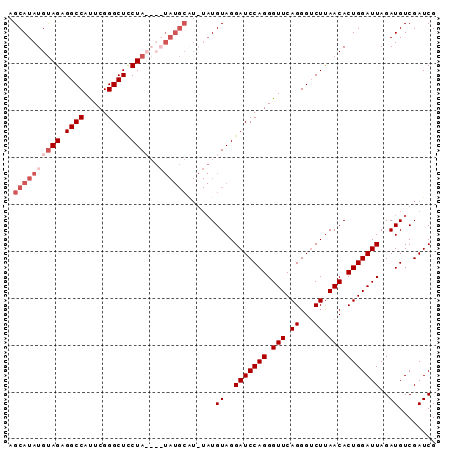
| Location | 10,867,295 – 10,867,385 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 87.80 |
| Mean single sequence MFE | -20.07 |
| Consensus MFE | -15.99 |
| Energy contribution | -17.68 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640120 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
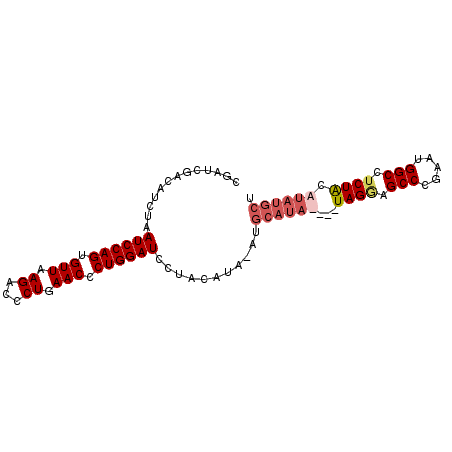
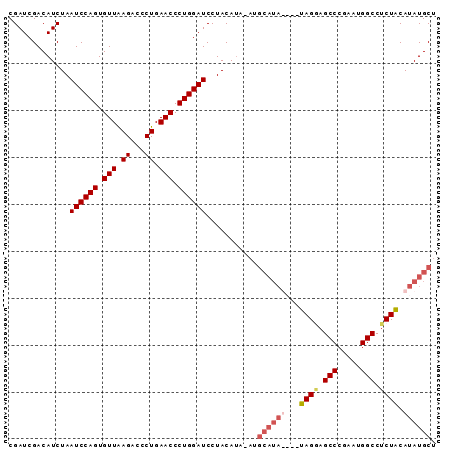
>2R_DroMel_CAF1 10867295 90 - 20766785 CGAUCGACAUCUAAUCCAGUGUUAAGACCCUGAACCCUGGAUCCUACAUA-ACGCAUAUACAUAGGAGCCCGAGUGGCCUCUACAUAUGCU .............((((((.(((.((...)).))).))))))........-..((((((...(((..(((.....)))..))).)))))). ( -21.40) >DroSec_CAF1 7825 87 - 1 CGAUCGACAUCUAAUCCAGUGUUAAGACCCUGAACCCUGGAUCCUACAUACAUGCAUA----UAGGAGCCCGAAUGGCCUCUACAUAUGCU .............((((((.(((.((...)).))).))))))...........(((((----(((..(((.....)))..)))..))))). ( -20.30) >DroSim_CAF1 8181 91 - 1 CGAUCGACAUCUAAUCCAGUGUUAAGACCCUGAACCCUGGAUCCUACAUACAUGCAUAUACAUAGAAGCCCGAAUGGCCUCUACAUAUGCU .............((((((.(((.((...)).))).))))))...........((((((...((((.(((.....))).)))).)))))). ( -20.90) >DroEre_CAF1 9147 77 - 1 CGAUCGACAUCUAAUCCAGUGUUGAGUCCCUGAACCCUGGAUCCUACA--------------UAGCAGCCCGAAUGGCCUCUGCAUAUGCU .............((((((.(((.((...)).))).))))))....((--------------((((((.((....))...)))).)))).. ( -17.70) >consensus CGAUCGACAUCUAAUCCAGUGUUAAGACCCUGAACCCUGGAUCCUACAUA_AUGCAUA____UAGGAGCCCGAAUGGCCUCUACAUAUGCU .............((((((.(((.((...)).))).))))))...........((((((...((((.(((.....))).)))).)))))). (-15.99 = -17.68 + 1.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:21 2006