
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,759,021 – 10,759,129 |
| Length | 108 |
| Max. P | 0.821518 |

| Location | 10,759,021 – 10,759,129 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.14 |
| Mean single sequence MFE | -28.29 |
| Consensus MFE | -12.92 |
| Energy contribution | -12.37 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.46 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.821518 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
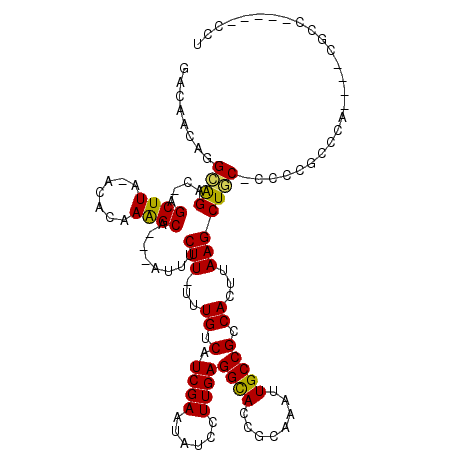
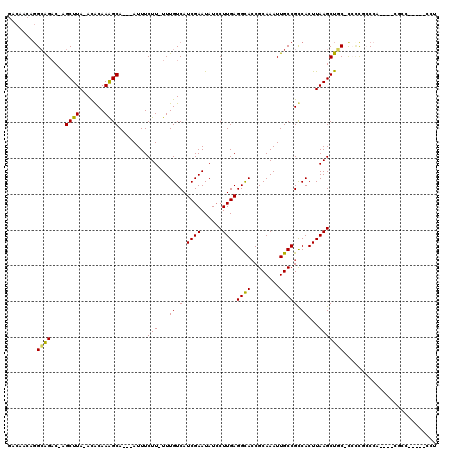
>2R_DroMel_CAF1 10759021 108 - 20766785 -----CUGGCAGAC-GGCUUA-AUACAAAGCA---AUUUCUU-UUUGUCAUCGAAUAUCCUUGAGGUACCGCAAAUUGCCGUCACUUAAGCUGC-CCACGCCCAACUGCGCCGAGCACCU -----..(((((..-.((((.-.....)))).---.......-................((((((..((.((.....)).))..))))))))))-)..........(((.....)))... ( -22.50) >DroVir_CAF1 29415 104 - 1 GACAACGGGCAGAC-AGCUUA-ACACAAGGCA---AUUUCUU-UUUGUCAUCGAAUAUCCUUGAGGCAACGCAAAUUGCCGCCACUUAAGCUGC-CCCCACCCA----CGCC-----CCU ......(((..(.(-((((((-(.....((((---((((...-.(((((.((((......)))))))))...)))))))).....)))))))))-.))).....----....-----... ( -30.70) >DroGri_CAF1 25568 104 - 1 GACAACAGGCAGAC-AGCUUA-ACGCAAAGCA---AUUUCUU-UUUGUCAUCGAAUAUCCUUGAGGCACCGCAAAUUGCCGCCACUUAAGCUGC-CCCCAACCA----CGCC-----UCC .......((..(.(-((((((-(.((...(((---((((...-..((((.((((......))))))))....))))))).))...)))))))))-..)).....----....-----... ( -23.50) >DroWil_CAF1 30554 117 - 1 GCCUGCUGGCAGACAGGCUUA-ACACAAAGCAAAAAUUUCUU-UUUGUCAUCGAAUAUGCUUGAGGCACCGCAAAUUGCCGUCACUUAAGCUCC-UGUUGGCCGCCUGCAGUUAUCCCCU ..((((.(((.((((((....-.......(((((((....))-)))))..........(((((((..((.((.....)).))..))))))).))-))))....))).))))......... ( -33.90) >DroMoj_CAF1 31166 105 - 1 GACAACAGGCAGAC-AGCUUA-ACGCAAGGCA---AUUUCUUUUUUGUCAUCGAAUAUCCUUGAGGCACCGCAAAUUGCCGCCACUUAAGCUGC-UGCUGCCGC----CACC-----ACA .......(((((.(-((((((-(.....((((---((((......((((.((((......))))))))....)))))))).....)))))))).-..)))))..----....-----... ( -34.20) >DroAna_CAF1 21232 92 - 1 --------GGGGCC-GGCUUAAACACAAAGCA---AUUUCUU-UUUGUCAUCGAAUAUCCUUGAGGCACCGCAAAUUGCCGUCACUUAAGCUCCACCCCGCCC---------------CU --------((((..-(((((((.......(((---((((...-..((((.((((......))))))))....)))))))......)))))))...))))....---------------.. ( -24.92) >consensus GACAACAGGCAGAC_AGCUUA_ACACAAAGCA___AUUUCUU_UUUGUCAUCGAAUAUCCUUGAGGCACCGCAAAUUGCCGCCACUUAAGCUGC_CCCCGCCCA____CGCC_____CCU ........((((....((((.......))))........(((...((.(.((((......))))((((........))))).))...))))))).......................... (-12.92 = -12.37 + -0.55)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:48:43 2006