| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,741,137 – 10,741,228 |
| Length | 91 |
| Max. P | 0.995727 |

| Location | 10,741,137 – 10,741,228 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 70.43 |
| Mean single sequence MFE | -28.82 |
| Consensus MFE | -14.80 |
| Energy contribution | -13.70 |
| Covariance contribution | -1.10 |
| Combinations/Pair | 1.47 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.51 |
| SVM decision value | 2.44 |
| SVM RNA-class probability | 0.993925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 10741137 91 + 20766785 UUCUUGGGACCAACGGCAACUUUUCGUGGCUUGUGCCA----------CGGGGUCUCUCACUCUCUC---UUGGCC-CCCUUCCCAUGUGCCCCUCCAUCAUAGC ......(((.....((((.......(((((....))))----------)((((((............---..))))-)).........))))..)))........ ( -27.24) >DroSim_CAF1 7826 95 + 1 UUCUUGGGGCCAACGGCAACUUUUCGUGGCUUGUGCCA----------CGGGGUCUCUCUCUCUCUCUCUUUGGCCCCCCUUUCCAUAAGCCCCUCCAUCAUAGC .....(((((....((.........(((((....))))----------)((((((.................)))))).....))....)))))........... ( -30.73) >DroAna_CAF1 7860 96 + 1 UUCUUGGGGCCAACGGCAACUUUUCGUGGCUUGUGCCGGAGCUCCAAACCGAAUCUACCUUCC--------UGCCU-CCAACCCCAUUUACCCGGGGAUCGCAGC ...(((((((...(((((.((......))....)))))..)))))))...............(--------(((..-....((((........))))...)))). ( -28.50) >consensus UUCUUGGGGCCAACGGCAACUUUUCGUGGCUUGUGCCA__________CGGGGUCUCUCUCUCUCUC___UUGGCC_CCCUUCCCAUAUGCCCCUCCAUCAUAGC ....(((((.....(((.........((((....))))...........(((........)))..........))).....)))))................... (-14.80 = -13.70 + -1.10)

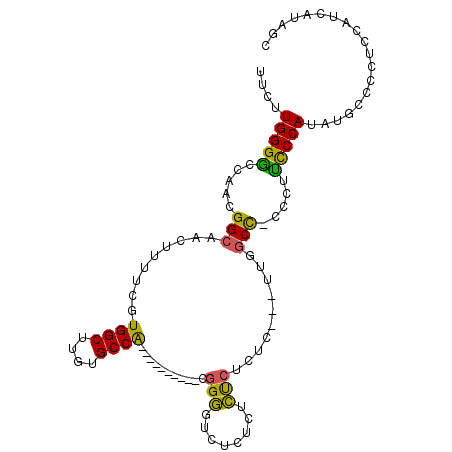

| Location | 10,741,137 – 10,741,228 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 70.43 |
| Mean single sequence MFE | -32.97 |
| Consensus MFE | -19.46 |
| Energy contribution | -18.25 |
| Covariance contribution | -1.21 |
| Combinations/Pair | 1.31 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.59 |
| SVM decision value | 2.61 |
| SVM RNA-class probability | 0.995727 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
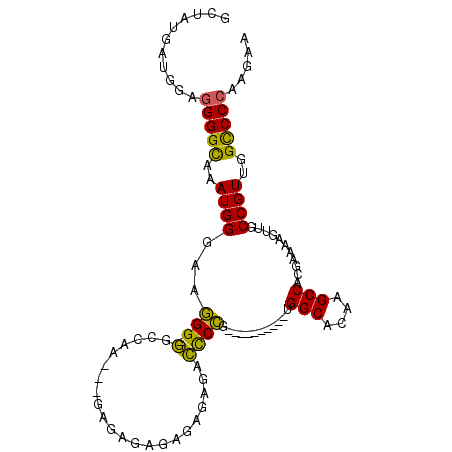
>2R_DroMel_CAF1 10741137 91 - 20766785 GCUAUGAUGGAGGGGCACAUGGGAAGGG-GGCCAA---GAGAGAGUGAGAGACCCCG----------UGGCACAAGCCACGAAAAGUUGCCGUUGGUCCCAAGAA ...........(((((..((((.(((((-(.(((.---.......)).)...))))(----------((((....)))))......)).))))..)))))..... ( -30.90) >DroSim_CAF1 7826 95 - 1 GCUAUGAUGGAGGGGCUUAUGGAAAGGGGGGCCAAAGAGAGAGAGAGAGAGACCCCG----------UGGCACAAGCCACGAAAAGUUGCCGUUGGCCCCAAGAA ...........((((((.((((...((((.......................))))(----------((((....))))).........)))).))))))..... ( -34.10) >DroAna_CAF1 7860 96 - 1 GCUGCGAUCCCCGGGUAAAUGGGGUUGG-AGGCA--------GGAAGGUAGAUUCGGUUUGGAGCUCCGGCACAAGCCACGAAAAGUUGCCGUUGGCCCCAAGAA ......(((....)))...((((((..(-.((((--------((((......)))((((((..((....)).))))))........))))).)..)))))).... ( -33.90) >consensus GCUAUGAUGGAGGGGCAAAUGGGAAGGG_GGCCAA___GAGAGAGAGAGAGACCCCG__________UGGCACAAGCCACGAAAAGUUGCCGUUGGCCCCAAGAA ...........(((((..((((...(((.(......................))))............(((....)))...........))))..)))))..... (-19.46 = -18.25 + -1.21)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:48:31 2006