
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,556,704 – 10,556,796 |
| Length | 92 |
| Max. P | 0.792390 |

| Location | 10,556,704 – 10,556,796 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 76.67 |
| Mean single sequence MFE | -15.28 |
| Consensus MFE | -8.92 |
| Energy contribution | -10.70 |
| Covariance contribution | 1.78 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.653416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
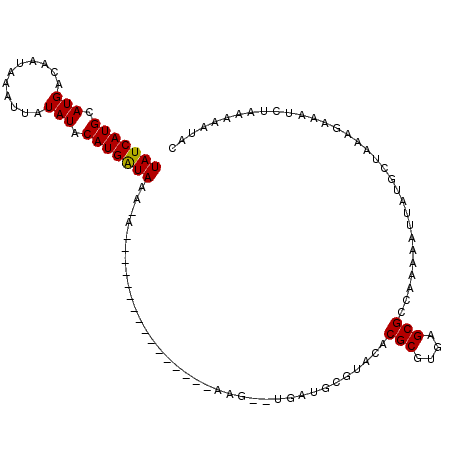
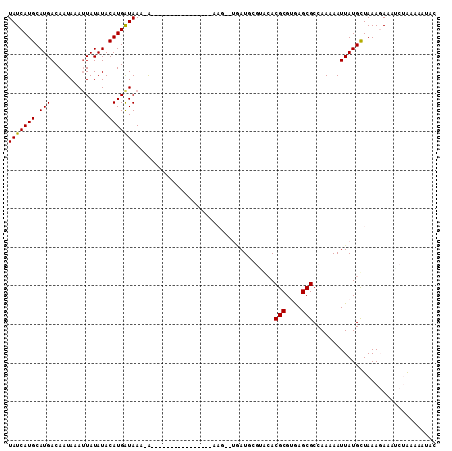
>2R_DroMel_CAF1 10556704 92 + 20766785 UAUCAUGCAUGACAAUA-AUUAUAUACAUGGUAAA-G----------------AAG-UUAAUGCAUGCACGCGUGAGCGCCAAAAAUUAUGCUAAAGAAUUCUAUAAAUAC ..((.((((((.((.((-(((...(((...)))..-.----------------.))-))).))))))))(((....))).................))............. ( -15.60) >DroPse_CAF1 38744 94 + 1 UAUCAUGCAUGACAAUAAAUUAUAUACAUGAUAAC-G----------------AAGGAUGAUCCGAGCACGCGUGAGCGCCAAAAAUUAUGCUAAAGAAAUCUACGAAUAC (((((((.(((...........))).))))))).(-(----------------..((((......(((((((....)))..........))))......)))).))..... ( -16.70) >DroGri_CAF1 32789 88 + 1 UAUCAUGCAUGACAAUAAAUUAUAUACAUGAUAAA-A----------------A------AUGCGUAAACGCGCAAGCGCCAAAAAUUAUGCUAAACAUAACAGAAAAGCU (((((((.(((...........))).)))))))..-.----------------.------..((.....(((....))).......(((((.....))))).......)). ( -14.00) >DroMoj_CAF1 40068 89 + 1 UAUCAUGCAUGACAAUAAAAUAUAUACAUGAUAAACA----------------U------AUGCGUACACGCGCAAGCGCCAAAAAUUAUGUUAAACAAAGCAGAAAAGCG (((((((.(((...........))).)))))))((((----------------(------(........(((....)))........)))))).......((......)). ( -16.49) >DroAna_CAF1 39754 104 + 1 UAUCAUGCAUGACAAUA-AUUAUAUACAUGGUAAA-AAAAAGAAACGAAAAACAAG-UUGAU----ACACGCGUGAGCGCCAAAAAUUAUGCUAAAGAACUCUAUAAAUAC ......((((((..(((-....)))...((((...-.......(((.........)-))...----....((....))))))....))))))................... ( -12.20) >DroPer_CAF1 39750 94 + 1 UAUCAUGCAUGACAAUAAAUUAUAUACAUGAUAAC-G----------------AAGGAUGAUCCGAGCACGCGUGAGCGCCAAAAAUUAUGCUAAAGAAAUCUACGAAUAC (((((((.(((...........))).))))))).(-(----------------..((((......(((((((....)))..........))))......)))).))..... ( -16.70) >consensus UAUCAUGCAUGACAAUAAAUUAUAUACAUGAUAAA_A________________AAG__UGAUGCGUACACGCGUGAGCGCCAAAAAUUAUGCUAAAGAAAUCUAAAAAUAC (((((((.(((...........))).)))))))....................................(((....)))................................ ( -8.92 = -10.70 + 1.78)


| Location | 10,556,704 – 10,556,796 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 76.67 |
| Mean single sequence MFE | -17.36 |
| Consensus MFE | -12.63 |
| Energy contribution | -13.13 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792390 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
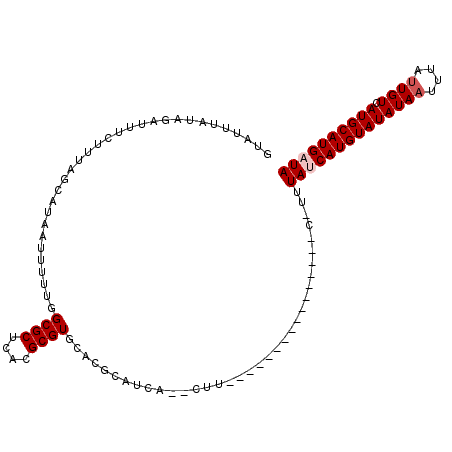
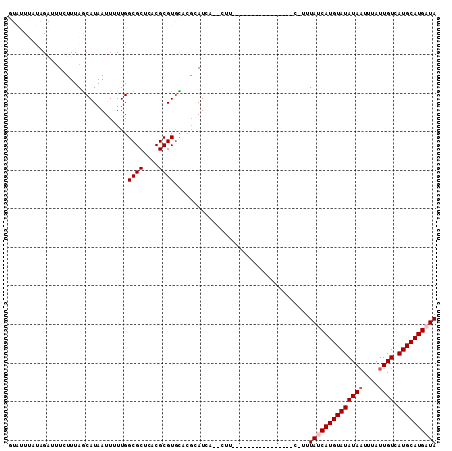
>2R_DroMel_CAF1 10556704 92 - 20766785 GUAUUUAUAGAAUUCUUUAGCAUAAUUUUUGGCGCUCACGCGUGCAUGCAUUAA-CUU----------------C-UUUACCAUGUAUAUAAU-UAUUGUCAUGCAUGAUA ...................((((........((((......)))))))).....-...----------------.-.....((((((((((..-...))).)))))))... ( -12.50) >DroPse_CAF1 38744 94 - 1 GUAUUCGUAGAUUUCUUUAGCAUAAUUUUUGGCGCUCACGCGUGCUCGGAUCAUCCUU----------------C-GUUAUCAUGUAUAUAAUUUAUUGUCAUGCAUGAUA .....((..(((.(((..(((((........(((....)))))))).)))..)))...----------------)-).((((((((((((((....)))).)))))))))) ( -17.60) >DroGri_CAF1 32789 88 - 1 AGCUUUUCUGUUAUGUUUAGCAUAAUUUUUGGCGCUUGCGCGUUUACGCAU------U----------------U-UUUAUCAUGUAUAUAAUUUAUUGUCAUGCAUGAUA .(((.....((((((.....))))))....)))......(((....)))..------.----------------.-..((((((((((((((....)))).)))))))))) ( -21.90) >DroMoj_CAF1 40068 89 - 1 CGCUUUUCUGCUUUGUUUAACAUAAUUUUUGGCGCUUGCGCGUGUACGCAU------A----------------UGUUUAUCAUGUAUAUAUUUUAUUGUCAUGCAUGAUA .((......)).......((((((.......((....))(((....))).)------)----------------))))(((((((((((((......))).)))))))))) ( -19.60) >DroAna_CAF1 39754 104 - 1 GUAUUUAUAGAGUUCUUUAGCAUAAUUUUUGGCGCUCACGCGUGU----AUCAA-CUUGUUUUUCGUUUCUUUUU-UUUACCAUGUAUAUAAU-UAUUGUCAUGCAUGAUA (((.....((((......((((.........((((....))))..----.....-..))))......))))....-..)))((((((((((..-...))).)))))))... ( -14.97) >DroPer_CAF1 39750 94 - 1 GUAUUCGUAGAUUUCUUUAGCAUAAUUUUUGGCGCUCACGCGUGCUCGGAUCAUCCUU----------------C-GUUAUCAUGUAUAUAAUUUAUUGUCAUGCAUGAUA .....((..(((.(((..(((((........(((....)))))))).)))..)))...----------------)-).((((((((((((((....)))).)))))))))) ( -17.60) >consensus GUAUUUAUAGAUUUCUUUAGCAUAAUUUUUGGCGCUCACGCGUGCACGCAUCA__CUU________________C_UUUAUCAUGUAUAUAAUUUAUUGUCAUGCAUGAUA ...............................((((....))))...................................((((((((((((((....)))).)))))))))) (-12.63 = -13.13 + 0.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:12 2006