
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,245,587 – 10,245,687 |
| Length | 100 |
| Max. P | 0.750270 |

| Location | 10,245,587 – 10,245,687 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.44 |
| Mean single sequence MFE | -18.75 |
| Consensus MFE | -11.23 |
| Energy contribution | -11.48 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.750270 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
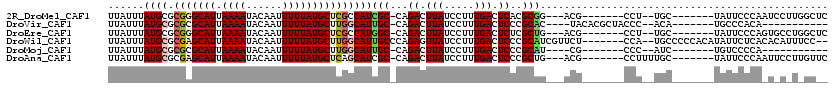
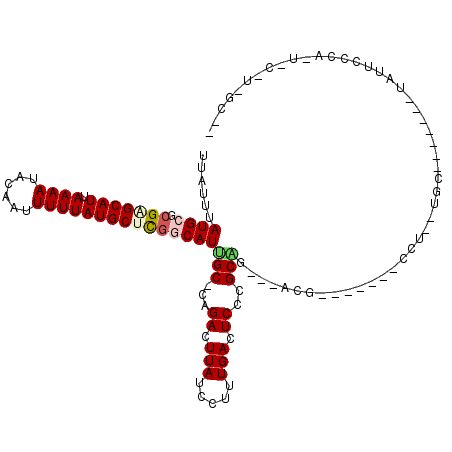
>2R_DroMel_CAF1 10245587 100 + 20766785 UUAUUUAUGCGCGGGCAUUAAAAUACAAUUUUUAUGCUCGCCAUCGC-CAGACUUAUCCUUUGACUCACGCGG---ACG-------CCU--UGC-------UAUUCCCAAUCCUUGGCUC ......(((.((((((((.((((......))))))))))))))).((-((((.(((.....))).))..((((---...-------..)--)))-------.............)))).. ( -22.70) >DroVir_CAF1 127925 95 + 1 UUAUUUAUGCGCGCGCAUUAAAAUACAAUUUUUAUGCUUGGCAUUGC-CAGACUUAUCCUUUGACUCCCGCAC----UACACGCUACCC--ACA-------UGCCCACA----------- .......((((...((((.((((......))))))))(((((...))-))).................)))).----.....((.....--...-------.)).....----------- ( -12.50) >DroEre_CAF1 78979 100 + 1 UUAUUUAUGCGCGGGCAUUAAAAUACAAUUUUUAUGCUCGCCAUGGC-CAGACUUAUCCUUUGACUCUCGCUG---ACG-------CCU--UGC-------UAUUCCCAGUGCCUGGCUC ......(((.((((((((.((((......)))))))))))))))(((-.(((.(((.....))).))).))).---..(-------((.--.((-------..........))..))).. ( -27.00) >DroWil_CAF1 90018 109 + 1 UUAUUUAUGCGCGAGCAUUAAAAUACAAUUUUUAUGCUUGGCAUUGCCCAGAGUUAUCCUUUGACUCCCGCAUCGUUCU-------CCA--UGCCCCCACAUAUUCUCACACAUUUCC-- ......((((.(((((((.((((......)))))))))))))))(((...((((((.....))))))..))).......-------...--...........................-- ( -21.20) >DroMoj_CAF1 129086 88 + 1 UUAUUUAUGCGCGCGCAUUAAAAUACAAUUUUUAUGCUUGGCAUUGC-CAGACUUAUCCUUUGACUCCCGCAU----CG-------CCC--AUC-------UGUCCCCA----------- ........(((.((((((.((((......))))))))(((((...))-)))..................))..----))-------)..--...-------........----------- ( -13.30) >DroAna_CAF1 76364 102 + 1 UUAUUUAUGCGCGAGCAUUAAAAUACAAUUUUUAUGCUCAGCAUCGC-CAGACUUAUCCUUUGACUCCCGCUG---ACG-------CCUUUUGC-------UAUUCCCAAUUCCUUGUUC ..........((((((((.((((......)))))))))).((.((((-..((.(((.....))).))..)).)---).)-------).....))-------................... ( -15.80) >consensus UUAUUUAUGCGCGAGCAUUAAAAUACAAUUUUUAUGCUCGGCAUUGC_CAGACUUAUCCUUUGACUCCCGCAG___ACG_______CCU__UGC_______UAUUCCCA_U_C_U_GC__ ......((((.(((((((.((((......)))))))))))))))(((...((.(((.....))).))..)))................................................ (-11.23 = -11.48 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:44:18 2006