
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 10,002,528 – 10,002,629 |
| Length | 101 |
| Max. P | 0.999307 |

| Location | 10,002,528 – 10,002,629 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 79.91 |
| Mean single sequence MFE | -19.86 |
| Consensus MFE | -13.08 |
| Energy contribution | -13.48 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.69 |
| Structure conservation index | 0.66 |
| SVM decision value | 3.50 |
| SVM RNA-class probability | 0.999307 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
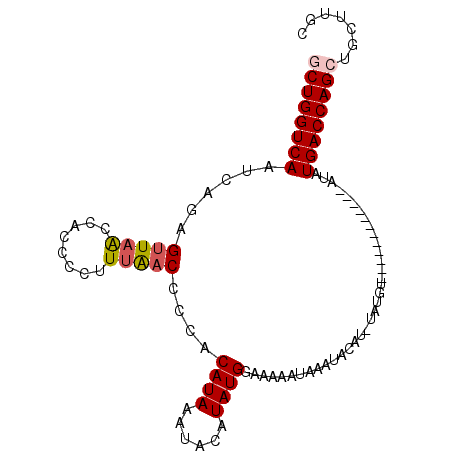
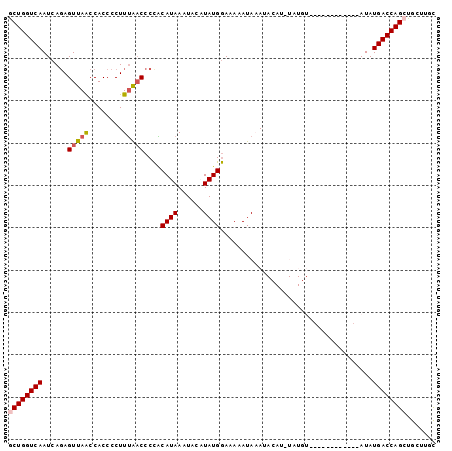
>2R_DroMel_CAF1 10002528 101 + 20766785 GCUGGUCAAUCACAGUUAACCACCCCUUUGACCCCACAUAAAUACAUAUGGAAAAAUAAAUACAUUUAUGUACAUACAUAUACAUAUGACCAGCUGCUUGC ((((((((.((...(((((........)))))....((((......))))))..............((((((........))))))))))))))....... ( -19.90) >DroSec_CAF1 5682 89 + 1 GCUGGUCAAUCAGAGUUAACCACCCCUUUGACCCCACAUAAAUACAUAUGGAAAAAUAAAUACAUAUAUGU------------AUAUGACCAGCUGCUUGU ((((((((.((((((..........))))))..........(((((((((..............)))))))------------)).))))))))....... ( -23.84) >DroSim_CAF1 6059 89 + 1 GCUGGUCAAUCAGAGUUAACCACCCCUUUAACCCCACAUAAAUACAUAUGGAAAAAUAAAUACAUAUAUGU------------AUAUGACCAGCUGCUUGC ((((((((......(((((........))))).........(((((((((..............)))))))------------)).))))))))....... ( -21.44) >DroEre_CAF1 5943 77 + 1 ACUGGUCAAUCAGAGUUAGCCACCCCUUUAACCCUGCAUAAAUA--UAUGAAAAAAUACAU----------------------AUAUGACCAGCUGCUCGC .((((((...(((.(((((........))))).))).....(((--((((........)))----------------------))))))))))........ ( -18.10) >DroYak_CAF1 5968 77 + 1 ACUGGUCAAUCAGAGUUAACCACCCCUUAACCCCCUCAUAAAUA--UAUGGAAAAAUACAU----------------------AUAUGACCAGCUGGUGGC .((((((.......(((((.......)))))..........(((--((((........)))----------------------))))))))))........ ( -16.00) >consensus GCUGGUCAAUCAGAGUUAACCACCCCUUUAACCCCACAUAAAUACAUAUGGAAAAAUAAAUACAU_UAUGU____________AUAUGACCAGCUGCUUGC ((((((((......(((((........)))))....((((......))))....................................))))))))....... (-13.08 = -13.48 + 0.40)

| Location | 10,002,528 – 10,002,629 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 79.91 |
| Mean single sequence MFE | -25.04 |
| Consensus MFE | -19.46 |
| Energy contribution | -20.66 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.993378 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
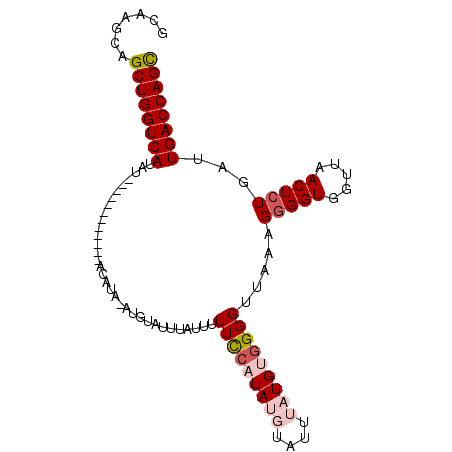
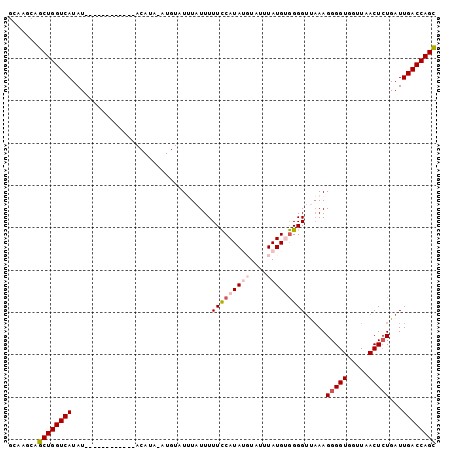
>2R_DroMel_CAF1 10002528 101 - 20766785 GCAAGCAGCUGGUCAUAUGUAUAUGUAUGUACAUAAAUGUAUUUAUUUUUCCAUAUGUAUUUAUGUGGGGUCAAAGGGGUGGUUAACUGUGAUUGACCAGC .......((((((((((((((((....))))))))..............((((((((....))))))))((((.((..(....)..)).)))))))))))) ( -28.90) >DroSec_CAF1 5682 89 - 1 ACAAGCAGCUGGUCAUAU------------ACAUAUAUGUAUUUAUUUUUCCAUAUGUAUUUAUGUGGGGUCAAAGGGGUGGUUAACUCUGAUUGACCAGC .......((((((((.((------------(((....)))))......(((((((((....))))))))).....(((((.....)))))...)))))))) ( -26.20) >DroSim_CAF1 6059 89 - 1 GCAAGCAGCUGGUCAUAU------------ACAUAUAUGUAUUUAUUUUUCCAUAUGUAUUUAUGUGGGGUUAAAGGGGUGGUUAACUCUGAUUGACCAGC .......((((((((..(------------(((....))))((((...(((((((((....))))))))).))))(((((.....)))))...)))))))) ( -26.40) >DroEre_CAF1 5943 77 - 1 GCGAGCAGCUGGUCAUAU----------------------AUGUAUUUUUUCAUA--UAUUUAUGCAGGGUUAAAGGGGUGGCUAACUCUGAUUGACCAGU .......(((((((((((----------------------(((........))))--))......((((((((..........))))))))..)))))))) ( -22.90) >DroYak_CAF1 5968 77 - 1 GCCACCAGCUGGUCAUAU----------------------AUGUAUUUUUCCAUA--UAUUUAUGAGGGGGUUAAGGGGUGGUUAACUCUGAUUGACCAGU .......(((((((((((----------------------(((........))))--))........((((((((.......))))))))...)))))))) ( -20.80) >consensus GCAAGCAGCUGGUCAUAU____________ACAUA_AUGUAUUUAUUUUUCCAUAUGUAUUUAUGUGGGGUUAAAGGGGUGGUUAACUCUGAUUGACCAGC .......((((((((.................................(((((((((....))))))))).....(((((.....)))))...)))))))) (-19.46 = -20.66 + 1.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:42:10 2006