
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,978,300 – 9,978,405 |
| Length | 105 |
| Max. P | 0.633180 |

| Location | 9,978,300 – 9,978,405 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 76.95 |
| Mean single sequence MFE | -36.15 |
| Consensus MFE | -24.46 |
| Energy contribution | -24.80 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.633180 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
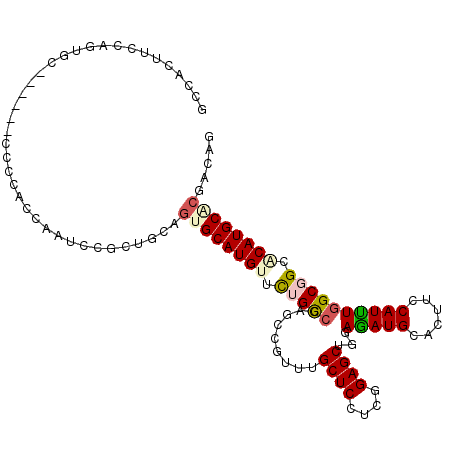
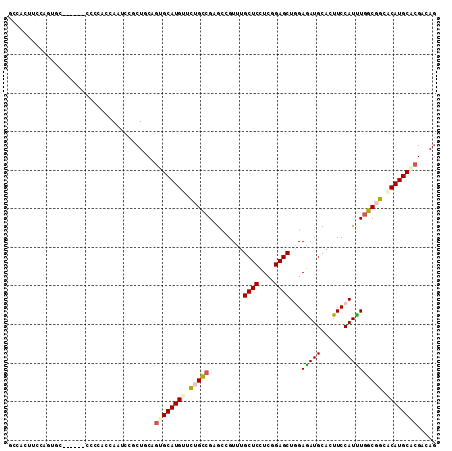
>2R_DroMel_CAF1 9978300 105 + 20766785 GCCACAUCUAGUGC------UCCCACUAAUCCGCUACAAUGCAUGUUCUGCCGAGCAGUUUGCUCUUCCGAGCUGGAAAUGCACUUCCAUUUGGCCGCACAUGCACGCCAG ((......(((((.------...)))))...........(((((((.(.((((((......((((....))))(((((......))))))))))).).))))))).))... ( -30.30) >DroPse_CAF1 49961 102 + 1 GGCAAC---AGCAG------UCCGGCCAAUCCGCUGCAGUGCAUGUUCUGCCGGGCCGUGUGCUCCUCUGAGCUGGAGAUGCACUUCCAUCUGGCGGCCCAUGCGCGACAG (....)---.((((------..(((.....))))))).(((((((..((((((((....((((..((((.....))))..)))).....))))))))..)))))))..... ( -44.20) >DroWil_CAF1 5077 111 + 1 UCAGUGUCAAGUGGUGUGAGCCCAACCAAUCCUCUCCAGUGCAUGUUUUGUCGUGCCGUUUGCUCCUCGGAGCUAGAGAUGCACUUCCAUUUAGCGGCACAUGCUCGACAA ....((((...((((.((....))))))..........(.(((((......)((((((((.((((....))))...(((((......))))))))))))))))).))))). ( -31.40) >DroYak_CAF1 37642 105 + 1 GCCACUUCUAAUGC------GCCCACCAAUCCGCUGCAGUGCAUGUUCUGCCGUGCAGUUUGCUCGUCCGAGCUGGAAAUGCACUUCCAUUUGGCCGCACAUGCACGUCAG ...........(((------((..........)).)))((((((((.(.((((((((.(..((((....))))..)...)))))........))).).))))))))..... ( -32.10) >DroMoj_CAF1 49253 105 + 1 GGUAAUCCCCCUGC------GCCGGCCAAUCCGCUGCAGUGCAUGUUCUGCCGCGCCGUCUGCUCCUCGGAGCUGGAGAUGCAUUUCCAUUUGGCGGCGCAUGCGCGACAG ..........((((------(.(((.....))).)))))....((((.(((.((((((((.((((....))))((((((....))))))...))))))))..))).)))). ( -43.00) >DroAna_CAF1 31990 105 + 1 GCCACUGCCAACAC------CCCCUCCAAUCCUCUACAGUGCAUGUUUUGCCGAGUGGUCUGCUCCUCGGAGCUCGAGAUGCACUUCCAUCUGGCGGCCCAUGCCCGGCAG (((((((.......------................))))(((((..((((((..(((..(((..((((.....))))..)))...)))..))))))..)))))..))).. ( -35.90) >consensus GCCACUUCCAGUGC______CCCCACCAAUCCGCUGCAGUGCAUGUUCUGCCGAGCCGUUUGCUCCUCGGAGCUGGAGAUGCACUUCCAUUUGGCGGCACAUGCACGACAG ......................................((((((((.(((((.........((((....))))...(((((......)))))))))).))))))))..... (-24.46 = -24.80 + 0.34)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:54 2006