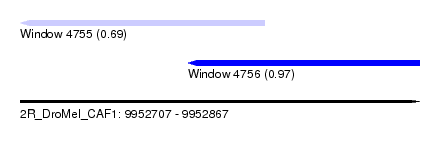
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,952,707 – 9,952,867 |
| Length | 160 |
| Max. P | 0.967609 |

| Location | 9,952,707 – 9,952,805 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 81.55 |
| Mean single sequence MFE | -28.72 |
| Consensus MFE | -20.88 |
| Energy contribution | -20.92 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686281 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
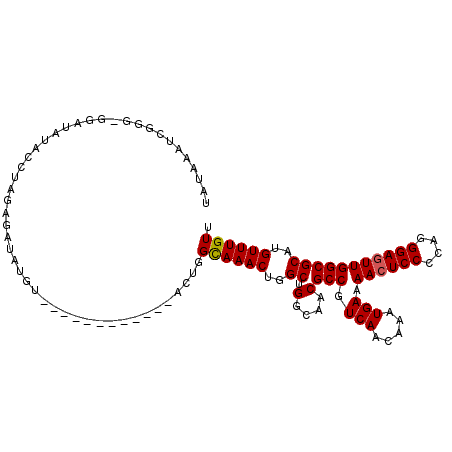
>2R_DroMel_CAF1 9952707 98 - 20766785 UAUAAAUCGGG-GAAUAUACCUAGAGAUAUGU------------AGUGGUAAACUGGCUGGCAACGCCGUCAACAAAUGAAAACUCCCCAGGGAGUUGGCGCAUGUUUGUU ........(((-.......)))..((((((((------------....((..((.(((.(....))))))..))....(..((((((....))))))..)))))))))... ( -26.10) >DroSec_CAF1 9941 98 - 1 UAUAAAUCGGG-GGAUAUUCCUAGAGAUAUGU------------ACUGGCAAACUGGCUGGCAACGCCGUCAACAAAUGAAAACUCCCCAGGGAGUUGGCGCAUGUUUGUU ((((.(((..(-((.....)))...))).)))------------)..(((((((..((.(....)(((.(((.....))).((((((....)))))))))))..))))))) ( -29.60) >DroSim_CAF1 10670 98 - 1 UAUAAAUCGGG-GGAUAUACCUAGAGAUAUGU------------ACUGGCAAACUGGCUGGCAACGCCGUCAACAAAUGAAAACUCCCCAGGGAGUUGGCGCAUGUUUGUU ((((.(((.((-(......)))...))).)))------------)..(((((((..((.(....)(((.(((.....))).((((((....)))))))))))..))))))) ( -29.70) >DroEre_CAF1 10072 110 - 1 CAUAAAUCGGG-GGAUAUACCUAGAGAUGUGUACAUACAUACAUACAGGCAAACUGGCUGGCAACGCCGUCAACAAAUGAAAACUCCCCAGGGAGUUGGCGCAUGUUUGUU ((((..((.((-(......))).))..)))).....(((.((((.(((.....)))((.(....)(((.(((.....))).((((((....))))))))))))))).))). ( -31.20) >DroAna_CAF1 8812 101 - 1 CAGAGCUUGUGCAGACAAACAA----------AAAUACAUAUAUACAUGCAAACUGGCUGGCAACGCCGUCAACAAAUGAAAAAUCCCCAGGGAGUUGGCGCAUGUUUGUU ....((....))..........----------........(((.((((((.....(((.(....))))((((((...(....).(((....))))))))))))))).))). ( -27.00) >consensus UAUAAAUCGGG_GGAUAUACCUAGAGAUAUGU____________ACUGGCAAACUGGCUGGCAACGCCGUCAACAAAUGAAAACUCCCCAGGGAGUUGGCGCAUGUUUGUU ................................................((((((..((.(....)(((.(((.....))).((((((....)))))))))))..)))))). (-20.88 = -20.92 + 0.04)

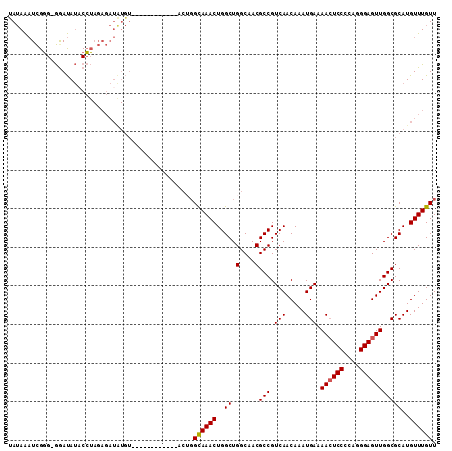
| Location | 9,952,774 – 9,952,867 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 85.08 |
| Mean single sequence MFE | -29.90 |
| Consensus MFE | -20.49 |
| Energy contribution | -23.27 |
| Covariance contribution | 2.78 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.77 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.967609 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
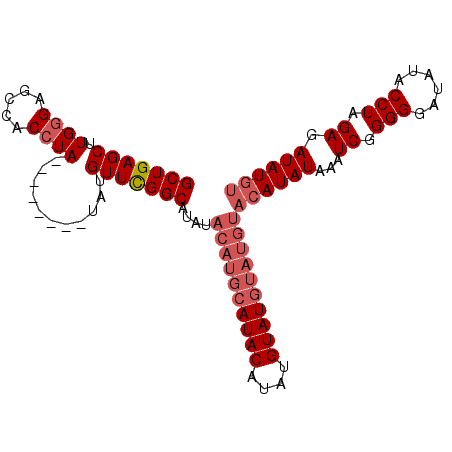
>2R_DroMel_CAF1 9952774 93 - 20766785 GCUGAGCUUGUGAGCCACCUACAUAUAUGUAUGUUCGGCAUAUACAUACAUACAUAUGUAU--------AUAUAAAUCGGGGAAUAUACCUAGAGAUAUGU (((((((..(((...))).((((....)))).)))))))(((((((((......)))))))--------))....(((.(((......)))...))).... ( -21.20) >DroSec_CAF1 10008 93 - 1 GCUGAGCUUGGGAGCCACCUA--------UAUGUUUGGCAUAUACAUGCAUACAUAUGUAUGUAUGUACAUAUAAAUCGGGGGAUAUUCCUAGAGAUAUGU ((..(((.((((.....))))--------...)))..))...(((((((((((....)))))))))))(((((...((..(((.....))).)).))))). ( -32.90) >DroSim_CAF1 10737 93 - 1 GCUGAGCUUGGGAGCCACCUA--------UAUGUUCGGCAUACACAUGCAUACAUAUGUAUGUAUGUACAUAUAAAUCGGGGGAUAUACCUAGAGAUAUGU (((((((.((((.....))))--------...)))))))....((((((((((....))))))))))((((((...((.(((......))).)).)))))) ( -35.60) >consensus GCUGAGCUUGGGAGCCACCUA________UAUGUUCGGCAUAUACAUGCAUACAUAUGUAUGUAUGUACAUAUAAAUCGGGGGAUAUACCUAGAGAUAUGU (((((((.((((.....))))...........)))))))....((((((((((....))))))))))((((((...((.(((......))).)).)))))) (-20.49 = -23.27 + 2.78)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:47 2006