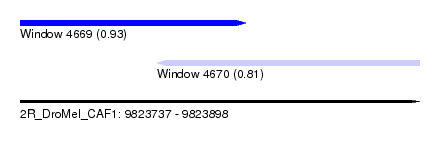
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,823,737 – 9,823,898 |
| Length | 161 |
| Max. P | 0.925028 |

| Location | 9,823,737 – 9,823,828 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 94.07 |
| Mean single sequence MFE | -22.97 |
| Consensus MFE | -20.14 |
| Energy contribution | -20.68 |
| Covariance contribution | 0.54 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925028 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
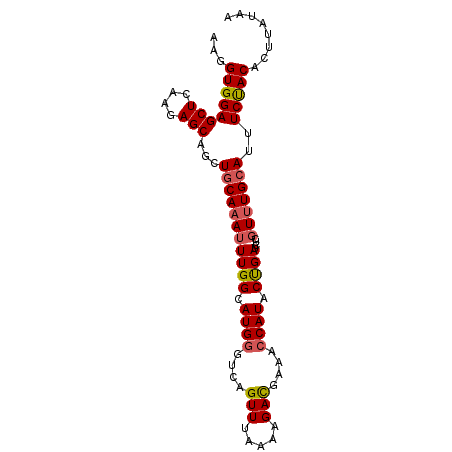
>2R_DroMel_CAF1 9823737 91 + 20766785 AAGGUGGAGCUCAAGAGCAGCUGCACAUUUGGCAUGGGUCAGUUUAAAAGACGAAACCAUACUGAUCUCGUUUGCAUUUCUACACUUAUAA ((((((.(((((....).)))).))).....(((.((((((((.................))))))))....))).........))).... ( -20.23) >DroSec_CAF1 61471 91 + 1 AAGGUGGAGCUCAAGAGCAGCUGCAAAUUUGGCAUGUGUCAGUUUAAAAGACGAAACCAUACUGAUCUCUUUUGCAUUUCUACACUUAUAA (((((((((((....)))...((((((.(..(.(((.((..(((.....)))...))))).)..).....))))))..))))).))).... ( -18.80) >DroSim_CAF1 63011 91 + 1 AAGGUGGAGCUCAAGAGCAGCUGCAAAUUUGGCAUGGGUCAGUUUAAAAGAUGAAACCAUACUGAUCUCGUUUGAAUUUCUACACUUAUAA (((((((((((....))).......((((..(.((((((((((.................))))))).))))..))))))))).))).... ( -20.23) >DroEre_CAF1 66201 91 + 1 AAGGUGGAGCUCAAGAGCAGCUGCAAAUUUGGCAUGGGUCAGUUUAAAAGACGAAACCAUACCGAUCUCGUUUGCAUUUCUACACCUUUAA ((((((..(((....)))...(((((((((((.((((....(((.....)))....)))).))))....)))))))......))))))... ( -26.60) >DroYak_CAF1 63297 91 + 1 AAGGUGGAGCUGAAAAGCAGCUGCAAAUUUGGCAUGGGUCAGUUUAGAAGACGAAACCAUACCGAUCUCGUUUGCAUUUCCACACCUUUAA ((((((.(((((.....)))))((((((((((.((((....(((.....)))....)))).))))....)))))).......))))))... ( -29.00) >consensus AAGGUGGAGCUCAAGAGCAGCUGCAAAUUUGGCAUGGGUCAGUUUAAAAGACGAAACCAUACUGAUCUCGUUUGCAUUUCUACACUUAUAA ...((((((((....)))...(((((((((((.((((....(((.....)))....)))).))))....)))))))..)))))........ (-20.14 = -20.68 + 0.54)


| Location | 9,823,792 – 9,823,898 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 84.81 |
| Mean single sequence MFE | -27.74 |
| Consensus MFE | -22.48 |
| Energy contribution | -23.00 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.812143 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
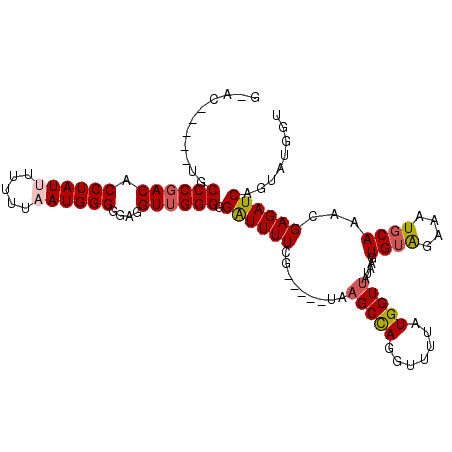
>2R_DroMel_CAF1 9823792 106 - 20766785 GUACCGACAUGCCCCACACCUAUUU-UUUAAUGGGGGAGGUUGGGGGAUUUUUG-----UAAGCCAGGUUUUAUGGUUAUAAGUGUAGAAAUGCAAACGAGAUCAGUAUGGU ..((((((...((((((.((((((.-...))))))....).)))))((((((.(-----(.(((((.......))))).....((((....)))).)))))))).)).)))) ( -29.90) >DroSec_CAF1 61526 98 - 1 ---------UGCCCGACACCUAUUUUUUUAAUGGGUGAGGUUGGGGGAUUUUAG-----UAAGCCAGGUUUUAUGGUUAUAAGUGUAGAAAUGCAAAAGAGAUCAGUAUGGU ---------..(((((((((((((.....)))))))...)))))).((((((..-----..(((((.......))))).....((((....))))...))))))........ ( -27.20) >DroSim_CAF1 63066 98 - 1 ---------UGCCCGACACCUAUUUUUUUAAUGGGGGAGGUUGGGGGAUUUUCG-----UAAGCCAGGUUUUAUGGUUAUAAGUGUAGAAAUUCAAACGAGAUCAGUAUGGU ---------..((((((.((((((.....))))))....)))))).(((((.((-----(.(((((.......))))).....((.(....).)).))))))))........ ( -24.90) >DroEre_CAF1 66256 104 - 1 GUAC-----UGCCCGACACCUAUUUUU---AUGGGGGAAGUUGGGGGGUUUUUAUUUUGUAAGCUAGGUUUUAUGGUUAAAGGUGUAGAAAUGCAAACGAGAUCGGUAUGGU ((((-----((((((((.(((((....---)))))....))))))........(((((((.(((((.......))))).....((((....)))).)))))))))))))... ( -27.10) >DroYak_CAF1 63352 99 - 1 GCAC-----UGCCCGACACCUAUUUUU---AUGGGGGAGGUUGGGGGAUUUUCA-----UACGCCAAGUUUUAUGGUUAAAGGUGUGGAAAUGCAAACGAGAUCGGUAUGGU ..((-----((((((((.(((((....---)))))....))))))(.(((((((-----(((((((.......)))).....)))))))))).).........))))..... ( -29.60) >consensus G_AC_____UGCCCGACACCUAUUUUUUUAAUGGGGGAGGUUGGGGGAUUUUCG_____UAAGCCAGGUUUUAUGGUUAUAAGUGUAGAAAUGCAAACGAGAUCAGUAUGGU ...........((((((.((((((.....))))))....)))))).((((((.........(((((.......))))).....((((....))))...))))))........ (-22.48 = -23.00 + 0.52)

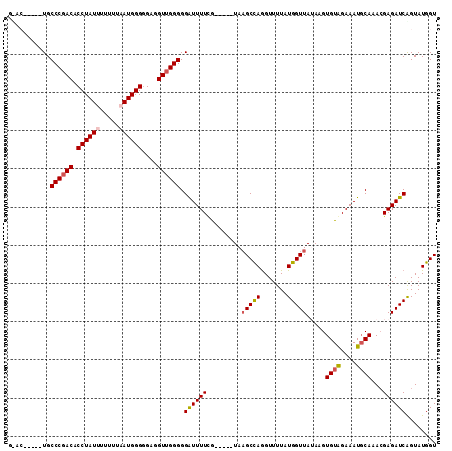
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:23 2006