
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,807,486 – 9,807,590 |
| Length | 104 |
| Max. P | 0.999941 |

| Location | 9,807,486 – 9,807,590 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.21 |
| Mean single sequence MFE | -21.36 |
| Consensus MFE | -10.91 |
| Energy contribution | -12.66 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.31 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.51 |
| SVM decision value | 3.74 |
| SVM RNA-class probability | 0.999578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
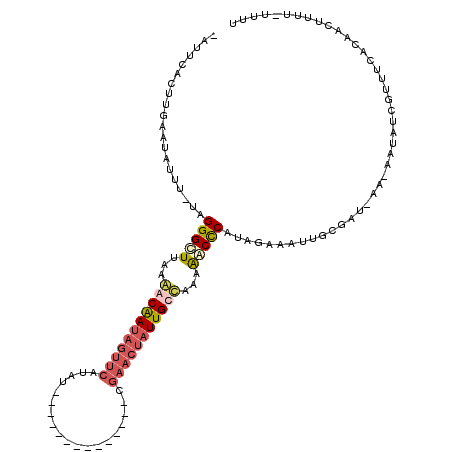
>2R_DroMel_CAF1 9807486 104 + 20766785 -AUUCACUUGAAUAUUU-UGGGGCUAAAAACAAUAGUUCAUAU--------------CGAACUAUUGUUAUAAACCCCUACAAAUUGCAUUGAAAAAUUUCUUUUCACAACUUUUUUUUU -.((((..((...((((-.((((.....(((((((((((....--------------.))))))))))).....))))...))))..)).)))).......................... ( -23.00) >DroSec_CAF1 45135 102 + 1 -AUACACUUCAGUAUUU-UUGGGCUGAAAACGAUAGUUCAUAU--------------CGAACUAUUGUUAAAAACCCAUAUAAAUUGCGUUUAA-AAUUUCUUUUCACAACUUUU-UUUU -..........((((((-.((((.....(((((((((((....--------------.))))))))))).....))))...))).)))......-....................-.... ( -19.00) >DroEre_CAF1 48706 117 + 1 UAUUCACUUGAAUUUGUUUAGGGCUUAAGACAAUAGUUCAUAUACCAUAUUUUACUAUGAACUAUUGCCUAAGGCUCAUGGAAUUCGCGAU-AA-AAUAUCGUUUCACAUCUUUU-UUUU .........((((((.....((((((.((.((((((((((((.............)))))))))))).)).))))))...))))))(((((-(.-..))))))............-.... ( -28.42) >DroYak_CAF1 47222 92 + 1 UAUUCACUUGAUUUUCCUUAGGGUUUA--ACAAUAGUUCAUA----------------------UUGCCUAAGACCCACGGAAAUUGCGAU-AA-AAUAUUGUUUCACACCUUU--UUUU ........(((.(((((...((((((.--.(((((.....))----------------------)))....))))))..)))))..(((((-(.-..)))))).))).......--.... ( -15.00) >consensus _AUUCACUUGAAUAUUU_UAGGGCUUAAAACAAUAGUUCAUAU______________CGAACUAUUGCCAAAAACCCAUAGAAAUUGCGAU_AA_AAUAUCGUUUCACAACUUUU_UUUU ....................(((((...(((((((((((...................)))))))))))...)))))........................................... (-10.91 = -12.66 + 1.75)


| Location | 9,807,486 – 9,807,590 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.21 |
| Mean single sequence MFE | -25.53 |
| Consensus MFE | -13.55 |
| Energy contribution | -16.30 |
| Covariance contribution | 2.75 |
| Combinations/Pair | 1.31 |
| Mean z-score | -3.77 |
| Structure conservation index | 0.53 |
| SVM decision value | 4.71 |
| SVM RNA-class probability | 0.999941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
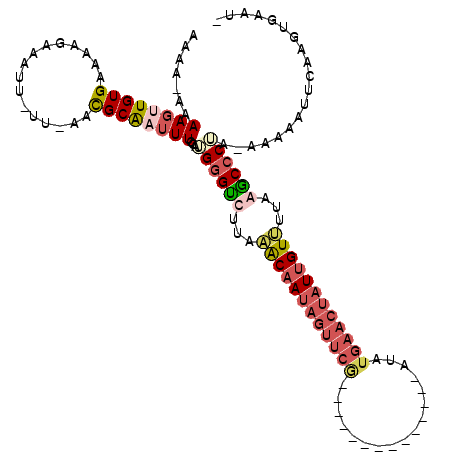
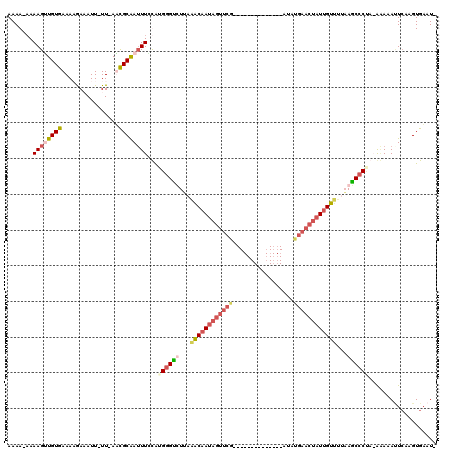
>2R_DroMel_CAF1 9807486 104 - 20766785 AAAAAAAAAGUUGUGAAAAGAAAUUUUUCAAUGCAAUUUGUAGGGGUUUAUAACAAUAGUUCG--------------AUAUGAACUAUUGUUUUUAGCCCCA-AAAUAUUCAAGUGAAU- .......((((((((((((....)))))....)))))))...((((((...(((((((((((.--------------....)))))))))))...)))))).-....((((....))))- ( -30.60) >DroSec_CAF1 45135 102 - 1 AAAA-AAAAGUUGUGAAAAGAAAUU-UUAAACGCAAUUUAUAUGGGUUUUUAACAAUAGUUCG--------------AUAUGAACUAUCGUUUUCAGCCCAA-AAAUACUGAAGUGUAU- ....-..((((((((((((....))-))...))))))))...((((((...(((.(((((((.--------------....))))))).)))...)))))).-..((((....))))..- ( -25.20) >DroEre_CAF1 48706 117 - 1 AAAA-AAAAGAUGUGAAACGAUAUU-UU-AUCGCGAAUUCCAUGAGCCUUAGGCAAUAGUUCAUAGUAAAAUAUGGUAUAUGAACUAUUGUCUUAAGCCCUAAACAAAUUCAAGUGAAUA ....-..(((((((......)))))-))-.(((((((((....(.((...(((((((((((((((.............)))))))))))))))...)).)......)))))..))))... ( -27.02) >DroYak_CAF1 47222 92 - 1 AAAA--AAAGGUGUGAAACAAUAUU-UU-AUCGCAAUUUCCGUGGGUCUUAGGCAA----------------------UAUGAACUAUUGU--UAAACCCUAAGGAAAAUCAAGUGAAUA ....--.(((((((......)))))-))-.((((..(((((..((((.((((.(((----------------------((.....))))))--)))))))...))))).....))))... ( -19.30) >consensus AAAA_AAAAGUUGUGAAAAGAAAUU_UU_AACGCAAUUUCCAUGGGUCUUAAACAAUAGUUCG______________AUAUGAACUAUUGUUUUAAGCCCUA_AAAAAUUCAAGUGAAU_ .......((((((((................))))))))...((((((...((((((((((((.................))))))))))))...))))))................... (-13.55 = -16.30 + 2.75)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:11 2006