| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,680,942 – 9,681,054 |
| Length | 112 |
| Max. P | 0.728644 |

| Location | 9,680,942 – 9,681,054 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 90.16 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -21.68 |
| Energy contribution | -21.72 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.728644 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 9680942 112 + 20766785 AAAUUGUUAAUUUGGCGACAGUCUGGCAACUCCGCUGCUACUAAGUUAUAUUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGAAGAGCAGGAGUAAUGCCAGCU ..((((((........))))))(((((((((((((((((....))).......((((((....((((((.......))))))..))))))...))).)))))..)))))).. ( -34.50) >DroSec_CAF1 3383 111 + 1 AAAUUGUUAAUUGCGCGACAGUCUGGCAACUCCGCUGCAACUAAGUUAUAAUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGAAGAGUA-UAGUAAUGCCAGCU ..((((((........))))))((((((((((.(((.......))).......((((((....((((((.......))))))..))))))..)))).-......)))))).. ( -26.41) >DroSim_CAF1 3374 111 + 1 AAAUUGUUAAUUGCGCGACAGUCUGGCAACUCCGCUGCAGCUAAGUUAUAUUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGAAUAGCA-UAGUAAUGCCAGCU ...((((((((((.....)))).))))))....((((((((((.(((((....((((((....((((((.......))))))..)))))).))))).-))))..)).)))). ( -30.30) >DroEre_CAF1 3454 105 + 1 ------UUAAUUUCGCGACAGUGUGGCAACUCCACUGCAUCUAAGUUAUAUUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGGAGAGCA-CAUUAGUGCCAGCU ------........(((.(((((.(....)..))))))............((.((((((....((((((.......))))))..)))))).)).(((-(....))))..)). ( -28.20) >DroYak_CAF1 3406 105 + 1 ------UUAAUUUCGCGACAGUGUGGCAACUCCACUGCAACUAAGUUAUAUUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGGAGAGCA-CAGUAGUGCCAGCU ------........(((.(((((.(....)..))))))............((.((((((....((((((.......))))))..)))))).)).(((-(....))))..)). ( -28.20) >consensus AAAUUGUUAAUUUCGCGACAGUCUGGCAACUCCGCUGCAACUAAGUUAUAUUGCUCGCCAAAUUUAUAUUAUAUUUGUAUAACGGGCGAGAAGAGCA_CAGUAAUGCCAGCU ...................(((.(((((.......(((...(((......)))((((((....((((((.......))))))..))))))....))).......)))))))) (-21.68 = -21.72 + 0.04)

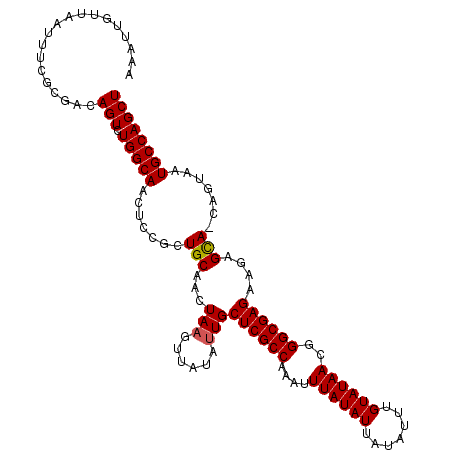
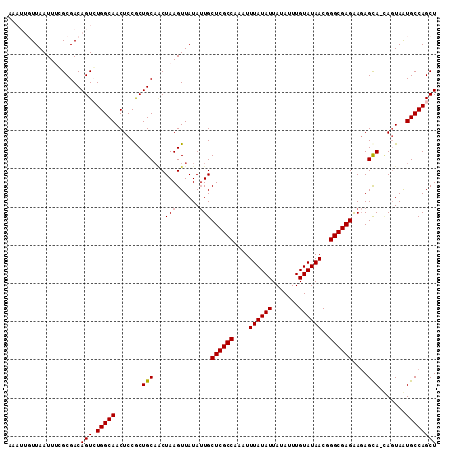
| Location | 9,680,942 – 9,681,054 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 90.16 |
| Mean single sequence MFE | -26.82 |
| Consensus MFE | -19.44 |
| Energy contribution | -19.68 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.578827 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 9680942 112 - 20766785 AGCUGGCAUUACUCCUGCUCUUCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAAUAUAACUUAGUAGCAGCGGAGUUGCCAGACUGUCGCCAAAUUAACAAUUU ((((((((..(((((((((((.((((((..(((((.........)))))....))))))...........)).))))..))))))))))).))................... ( -29.70) >DroSec_CAF1 3383 111 - 1 AGCUGGCAUUACUA-UACUCUUCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAUUAUAACUUAGUUGCAGCGGAGUUGCCAGACUGUCGCGCAAUUAACAAUUU ..............-.......((((((..(((((.........)))))....)))))).........((((((((.((((((((....)))).))))))))))))...... ( -24.70) >DroSim_CAF1 3374 111 - 1 AGCUGGCAUUACUA-UGCUAUUCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAAUAUAACUUAGCUGCAGCGGAGUUGCCAGACUGUCGCGCAAUUAACAAUUU (((((((((....)-)))((((((((((..(((((.........)))))....)))))).)))).....)))))((.((((((((....)))).))))))............ ( -28.40) >DroEre_CAF1 3454 105 - 1 AGCUGGCACUAAUG-UGCUCUCCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAAUAUAACUUAGAUGCAGUGGAGUUGCCACACUGUCGCGAAAUUAA------ .((.(((((....)-))))...((((((..(((((.........)))))....))))))...............((((((.........)))))).))........------ ( -24.60) >DroYak_CAF1 3406 105 - 1 AGCUGGCACUACUG-UGCUCUCCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAAUAUAACUUAGUUGCAGUGGAGUUGCCACACUGUCGCGAAAUUAA------ .((.((((....((-(((.(((((((((..(((((.........)))))....)))))(((((........)))))...))))..).)))).))))))........------ ( -26.70) >consensus AGCUGGCAUUACUA_UGCUCUUCUCGCCCGUUAUACAAAUAUAAUAUAAAUUUGGCGAGCAAUAUAACUUAGUUGCAGCGGAGUUGCCAGACUGUCGCGAAAUUAACAAUUU ..(((((...............((((((..(((((.........)))))....))))))......(((((.((....)).))))))))))...................... (-19.44 = -19.68 + 0.24)


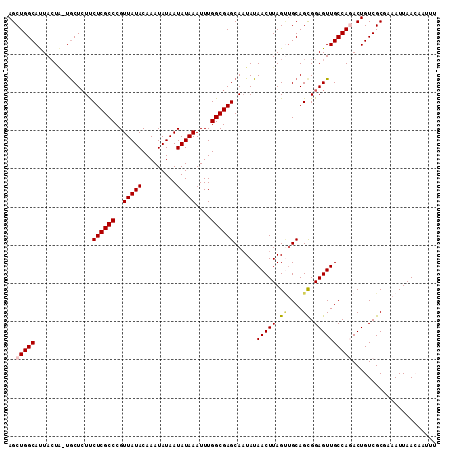
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:38:51 2006