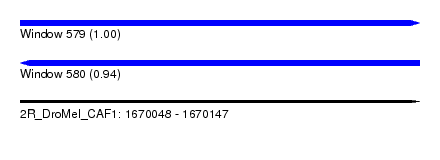
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 1,670,048 – 1,670,147 |
| Length | 99 |
| Max. P | 0.998986 |

| Location | 1,670,048 – 1,670,147 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.29 |
| Mean single sequence MFE | -34.08 |
| Consensus MFE | -30.59 |
| Energy contribution | -30.40 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.31 |
| SVM RNA-class probability | 0.998986 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
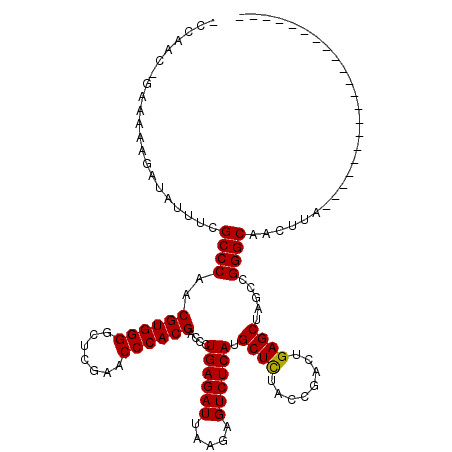
>2R_DroMel_CAF1 1670048 99 + 20766785 ACUAAUAUAAAAAUAUAACACGCCCAACGUGGGGCUCGAACCCACGACCCUGAGAUUAAGAGUCUCAUGCUCUACCGACUGAGCUAGCCGGGCCUCUUG--------------------- .....................((((..((((((.......))))))....((((((.....)))))).((((........)))).....))))......--------------------- ( -30.60) >DroPse_CAF1 86549 118 + 1 -CCGAC-GAAAAAGAUAUUUCGCCCAACGUGGGGCUCGAACCCACGACCCUGAGAUUAAGAGUCUCAUGCUCUACCGACUGAGCUAGCCGGGCAAUGUAGCCAGGGCUGGCCAAUAGAUC -.((..-((((......))))((....((((((.......))))))....((((((.....)))))).)).....))..((.(((((((.(((......)))..)))))))))....... ( -39.10) >DroSec_CAF1 55515 86 + 1 -------------NNNNNNNNGCCCAACGUGGGGCUCGAACCCACGACCCUGAGAUUAUGAGUCUCAUGCUUUAUCGACUGAGCUAGCCGGGCCACUUA--------------------- -------------........((((..((((((.......))))))....((((((.....)))))).((((........)))).....))))......--------------------- ( -28.00) >DroPer_CAF1 80509 118 + 1 -CCAAC-GAAAAAGAUAUUUCGCCCAACGUGGGGCUCGAACCCACGACCCUGAGAUUAAGAGUCUCAUGCUCUACCGACUGAGCUAGCCGGGCAAUAUAGCCAGGGUUGGCCAAUAGAUC -(((((-((((......))))((((..((((((.......))))))....((((((.....)))))).((((........)))).....))))............))))).......... ( -38.60) >consensus _CCAAC_GAAAAAGAUAUUUCGCCCAACGUGGGGCUCGAACCCACGACCCUGAGAUUAAGAGUCUCAUGCUCUACCGACUGAGCUAGCCGGGCAACUUA_____________________ .....................((((..((((((.......))))))....((((((.....)))))).((((........)))).....))))........................... (-30.59 = -30.40 + -0.19)

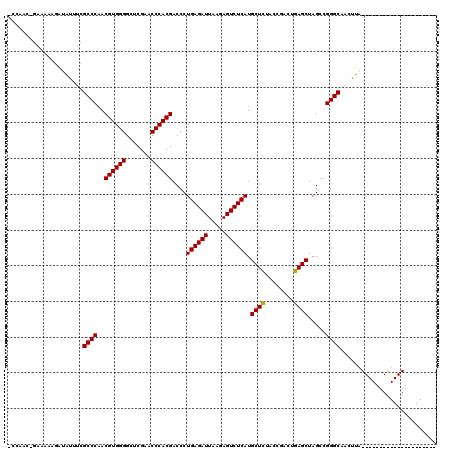
| Location | 1,670,048 – 1,670,147 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.29 |
| Mean single sequence MFE | -35.88 |
| Consensus MFE | -30.25 |
| Energy contribution | -30.75 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939409 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
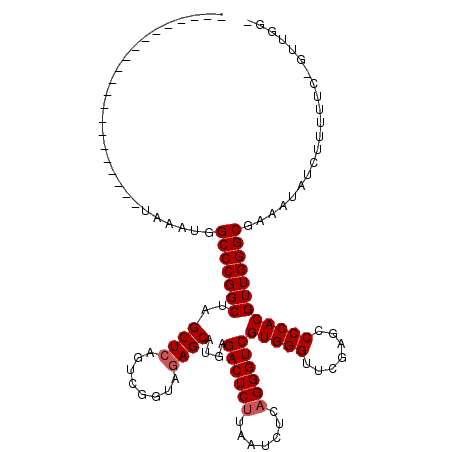
>2R_DroMel_CAF1 1670048 99 - 20766785 ---------------------CAAGAGGCCCGGCUAGCUCAGUCGGUAGAGCAUGAGACUCUUAAUCUCAGGGUCGUGGGUUCGAGCCCCACGUUGGGCGUGUUAUAUUUUUAUAUUAGU ---------------------.((.(.(((((((..((((........))))....((((((.......))))))(((((.......)))))))))))).).))................ ( -33.80) >DroPse_CAF1 86549 118 - 1 GAUCUAUUGGCCAGCCCUGGCUACAUUGCCCGGCUAGCUCAGUCGGUAGAGCAUGAGACUCUUAAUCUCAGGGUCGUGGGUUCGAGCCCCACGUUGGGCGAAAUAUCUUUUUC-GUCGG- (((....((((((....))))))..(((((((((..((((........))))....((((((.......))))))(((((.......))))))))))))))............-)))..- ( -42.80) >DroSec_CAF1 55515 86 - 1 ---------------------UAAGUGGCCCGGCUAGCUCAGUCGAUAAAGCAUGAGACUCAUAAUCUCAGGGUCGUGGGUUCGAGCCCCACGUUGGGCNNNNNNNN------------- ---------------------......(((((((..(((..........)))....(((((.........)))))(((((.......))))))))))))........------------- ( -25.20) >DroPer_CAF1 80509 118 - 1 GAUCUAUUGGCCAACCCUGGCUAUAUUGCCCGGCUAGCUCAGUCGGUAGAGCAUGAGACUCUUAAUCUCAGGGUCGUGGGUUCGAGCCCCACGUUGGGCGAAAUAUCUUUUUC-GUUGG- .......((((((....))))))....(((((((..((((........))))....((((((.......))))))(((((.......))))))))))))((((......))))-.....- ( -41.70) >consensus _____________________UAAAUGGCCCGGCUAGCUCAGUCGGUAGAGCAUGAGACUCUUAAUCUCAGGGUCGUGGGUUCGAGCCCCACGUUGGGCGAAAUAUCUUUUUC_GUUGG_ ...........................(((((((..((((........))))....((((((.......))))))(((((.......))))))))))))..................... (-30.25 = -30.75 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:34:16 2006