
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,513,981 – 9,514,126 |
| Length | 145 |
| Max. P | 0.998008 |

| Location | 9,513,981 – 9,514,101 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.50 |
| Mean single sequence MFE | -28.39 |
| Consensus MFE | -22.88 |
| Energy contribution | -23.68 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.815706 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
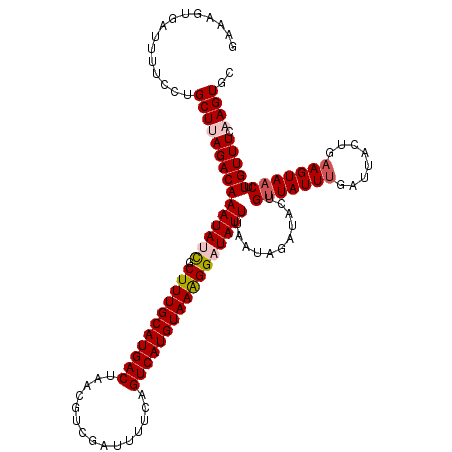
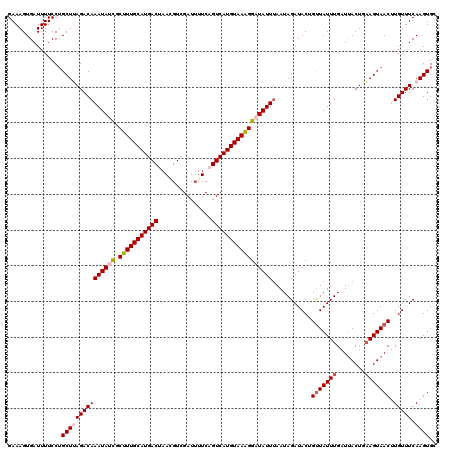
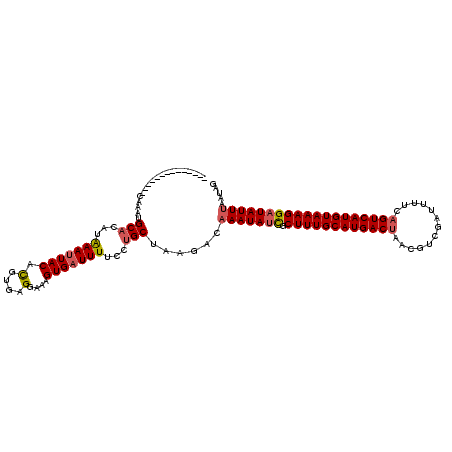
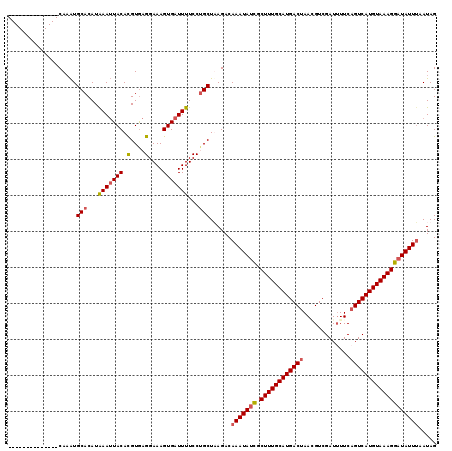
>2R_DroMel_CAF1 9513981 120 - 20766785 GAAAGUGAUUUUCCUGCUUAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUAAAUAGAUACUGUUAUUUGAUUACUGAAGUAACUUGUUUCAAGUGC ...............(((((((((((((((.((((((((((((.............))))))))))))))))))...........(((((((........))))))).))))).)))).. ( -28.82) >DroSec_CAF1 11006 120 - 1 GAAAGUGAUUUUCCAGCUAAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUUAAUAGAUACUGUUAUUUGAUUACUCAAGUAACUUGUUUCAAGUGC ...............(((.....(((((((.((((((((((((.............)))))))))))))))))))....((.((.(((((((((....)))))))))..)).)).))).. ( -31.92) >DroSim_CAF1 11051 120 - 1 GAAAGUGAUUUUCCUGCUAAGACAAAUAUCGCUUUGCAUGACUAAAGUCGAUGUUCAGUCAUGUAAAGGAUAUUUAAUAGAUACUGUUAUUUGAUUACUCUAGUAACUUGUUUCAAGUGC ...............(((.....(((((((.((((((((((((.............)))))))))))))))))))....((.((.((((((.((....)).))))))..)).)).))).. ( -27.72) >DroEre_CAF1 11240 120 - 1 GAAAGUGAUUUUCCUGCUUAGACAAAUAAUGCUUUGCAUGACUAAGGUCGAUUUUCCGUCAUGUAAAGGAUAUUUAAUAGAUACGGCUAUUUGCUAACUGAAGUAACGUGUUUCAAGUGG .............(..(((....(((((.(.(((((((((((...((........))))))))))))).))))))...(((((((..(((((........))))).))))))).)))..) ( -26.50) >DroYak_CAF1 11127 120 - 1 GAAAGUGAUUUUCCUGCUUAGACAAAUAAUGCUUUGCAUGACUAAGGUCGAUUUUCCGUCAUGUAAGGGAUAUUUAAUAGAUACGGUUAUUUGCUUACUGAAGUAACGUGUUUCAAGUGG .............(..((((((((....((.(((((((((((...((........))))))))))))).))..............(((((((........))))))).))))).)))..) ( -27.00) >consensus GAAAGUGAUUUUCCUGCUUAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUUAAUAGAUACUGUUAUUUGAUUACUGAAGUAACUUGUUUCAAGUGC ...............(((((((((((((((.(((((((((((...............)))))))))))))))))...........(((((((........))))))).))))).)))).. (-22.88 = -23.68 + 0.80)

| Location | 9,514,021 – 9,514,126 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 86.61 |
| Mean single sequence MFE | -27.55 |
| Consensus MFE | -24.58 |
| Energy contribution | -25.20 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.36 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.98 |
| SVM RNA-class probability | 0.998008 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
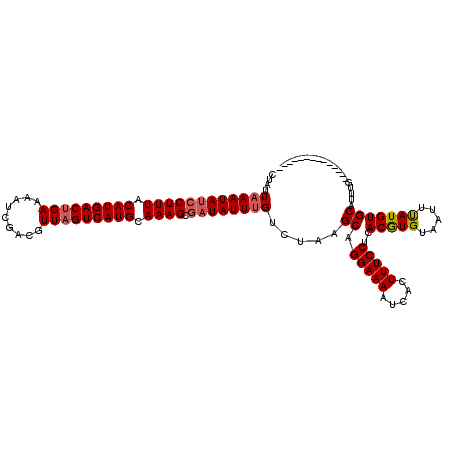
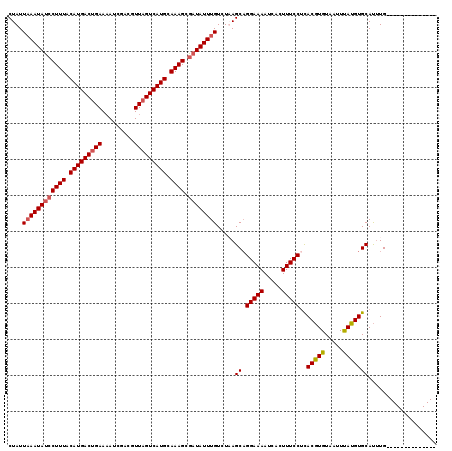
>2R_DroMel_CAF1 9514021 105 + 20766785 CUAUUUAAUAUCCUUUACAUGACUGAAAAUCGACGUUAGUCAUGCAAAGCGAUAUUUGUCUAAGCAGGAAAAUCACUUUCCUCACGUGUAAUUUAUGUGCAUUUG-------------- ......((((((((((.(((((((((.........))))))))).)))).)))))).......((((((((.....)))))).(((((.....))))))).....-------------- ( -25.90) >DroSec_CAF1 11046 105 + 1 CUAUUAAAUAUCCUUUACAUGACUGAAAAUCGACGUUAGUCAUGCAAAGCGAUAUUUGUCUUAGCUGGAAAAUCACUUUCCUCACGUGUAAUUUAUGUGCAUUUG-------------- (((.((((((((((((.(((((((((.........))))))))).)))).))))))))...)))..(((((.....))))).((((((.....))))))......-------------- ( -28.00) >DroSim_CAF1 11091 105 + 1 CUAUUAAAUAUCCUUUACAUGACUGAACAUCGACUUUAGUCAUGCAAAGCGAUAUUUGUCUUAGCAGGAAAAUCACUUUCCUCACGUGUAAUUUAUGUGCAUUUG-------------- (((.((((((((((((.((((((((((.......)))))))))).)))).))))))))...))).((((((.....))))))((((((.....))))))......-------------- ( -30.10) >DroEre_CAF1 11280 118 + 1 CUAUUAAAUAUCCUUUACAUGACGGAAAAUCGACCUUAGUCAUGCAAAGCAUUAUUUGUCUAAGCAGGAAAAUCACUUUCCAUACAUGUA-UUCAUGUGCACAUGUACAUAUGUAUGUA ....((((((..((((.((((((((........))...)))))).))))...)))))).....((.(((((.....)))))((((((((.-..((((....))))...)))))))))). ( -26.20) >consensus CUAUUAAAUAUCCUUUACAUGACUGAAAAUCGACGUUAGUCAUGCAAAGCGAUAUUUGUCUAAGCAGGAAAAUCACUUUCCUCACGUGUAAUUUAUGUGCAUUUG______________ ....((((((((((((.(((((((((.........))))))))).)))).)))))))).....((.(((((.....)))))..(((((.....)))))))................... (-24.58 = -25.20 + 0.62)

| Location | 9,514,021 – 9,514,126 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 86.61 |
| Mean single sequence MFE | -28.79 |
| Consensus MFE | -23.96 |
| Energy contribution | -24.64 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.25 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.56 |
| SVM RNA-class probability | 0.995321 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 9514021 105 - 20766785 --------------CAAAUGCACAUAAAUUACACGUGAGGAAAGUGAUUUUCCUGCUUAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUAAAUAG --------------.....(((...(((((((.(....)....)))))))...))).......((((((.((((((((((((.............))))))))))))))))))...... ( -28.52) >DroSec_CAF1 11046 105 - 1 --------------CAAAUGCACAUAAAUUACACGUGAGGAAAGUGAUUUUCCAGCUAAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUUAAUAG --------------.....((....(((((((.(....)....)))))))....))......(((((((.((((((((((((.............)))))))))))))))))))..... ( -27.72) >DroSim_CAF1 11091 105 - 1 --------------CAAAUGCACAUAAAUUACACGUGAGGAAAGUGAUUUUCCUGCUAAGACAAAUAUCGCUUUGCAUGACUAAAGUCGAUGUUCAGUCAUGUAAAGGAUAUUUAAUAG --------------.....(((...(((((((.(....)....)))))))...)))......(((((((.((((((((((((.............)))))))))))))))))))..... ( -29.42) >DroEre_CAF1 11280 118 - 1 UACAUACAUAUGUACAUGUGCACAUGAA-UACAUGUAUGGAAAGUGAUUUUCCUGCUUAGACAAAUAAUGCUUUGCAUGACUAAGGUCGAUUUUCCGUCAUGUAAAGGAUAUUUAAUAG ..(((((((.(((.((((....)))).)-)).)))))))..((((.........))))....(((((.(.(((((((((((...((........))))))))))))).))))))..... ( -29.50) >consensus ______________CAAAUGCACAUAAAUUACACGUGAGGAAAGUGAUUUUCCUGCUAAGACAAAUAUCGCUUUGCAUGACUAACGUCGAUUUUCAGUCAUGUAAAGGAUAUUUAAUAG ...................(((...(((((((.(....)....)))))))...)))......(((((((.((((((((((((.............)))))))))))))))))))..... (-23.96 = -24.64 + 0.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:12 2006