
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,377,441 – 9,377,534 |
| Length | 93 |
| Max. P | 0.962712 |

| Location | 9,377,441 – 9,377,534 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 71.63 |
| Mean single sequence MFE | -18.06 |
| Consensus MFE | -11.44 |
| Energy contribution | -11.40 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824100 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
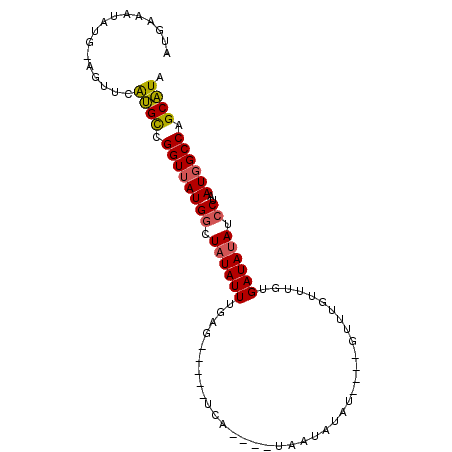
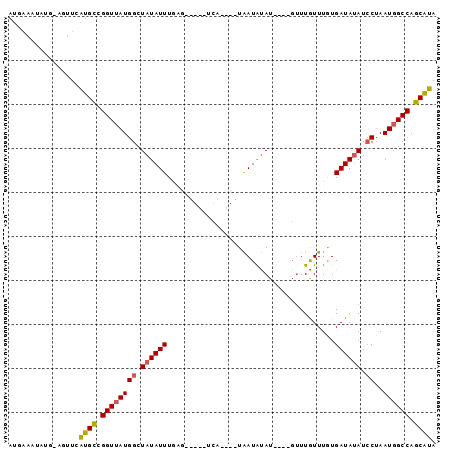
>2R_DroMel_CAF1 9377441 93 + 20766785 AUUAAUUAUGCAGUUCAUGCCGGUUAUGGCUAUAUUUCAGUUGUAUCAUGCAUAAUAUAU----GUUUGUUUGUGAUAUAUCCUAAUGGCCAGCAUA ......(((((.((....))......(((((((........(((((((((.((((.....----..)))).))))))))).....)))))))))))) ( -19.42) >DroSec_CAF1 2571 81 + 1 AUAAAAUAUGUAAUUUAUGCCGGUUAUGGCUAUAUUUGAG------------UAAUAUAU----GUUUGUUUGUGAUAGAUCCUAAUGGCCAGCAUA ...............(((((.((((((((...((((....------------.((((...----...))))...))))...))..)))))).))))) ( -13.40) >DroSim_CAF1 2403 93 + 1 AUGAAAUAUGUGUUUCAUGCCGGUUAUGGCUAUAUUUGAGUUGUAUCAUCCAUAAUAUAU----GUUUGUUUGUGAUAUAUCCUAAUGGCCAGCAUA (((((((....)))))))........(((((((....(.(.((((((((..((((.....----..))))..)))))))).))..)))))))..... ( -19.70) >DroEre_CAF1 2599 73 + 1 GUGAAAU----AGUUCACGUAGGUUAUGGCUAUAUUU---------------UGAUAUAUG-----UUGUUUAUGAUAUAUCCAAAUGGCCAGCGUA (((((..----..)))))((.((((((((.((((((.---------------.((((....-----.))))...)))))).)))..))))).))... ( -18.20) >DroYak_CAF1 2606 77 + 1 GU---------CGUUAGUGCCGGUUAUGGCUAUAUUUG-------UCA----UGAUGCAUGUAUGUUUAUUUAUGAUAUAUGCUAAUCGCCAGCAUA ..---------.....((((.(((((((((.......)-------)))----))..((((((((...........)))))))).....))).)))). ( -19.60) >consensus AUGAAAUAUG_AGUUCAUGCCGGUUAUGGCUAUAUUUGAG_____UCA____UAAUAUAU____GUUUGUUUGUGAUAUAUCCUAAUGGCCAGCAUA ................((((.((((((((.((((((......................................)))))).))..)))))).)))). (-11.44 = -11.40 + -0.04)

| Location | 9,377,441 – 9,377,534 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 71.63 |
| Mean single sequence MFE | -13.14 |
| Consensus MFE | -6.76 |
| Energy contribution | -7.52 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.51 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962712 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
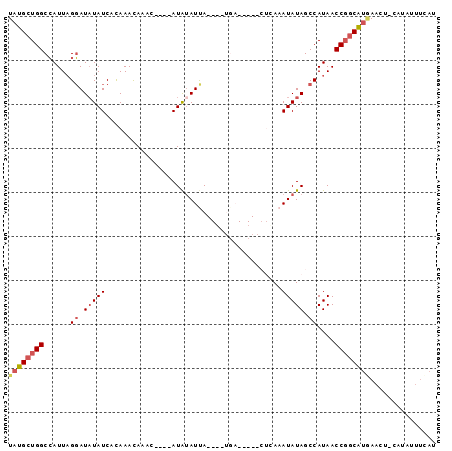
>2R_DroMel_CAF1 9377441 93 - 20766785 UAUGCUGGCCAUUAGGAUAUAUCACAAACAAAC----AUAUAUUAUGCAUGAUACAACUGAAAUAUAGCCAUAACCGGCAUGAACUGCAUAAUUAAU (((((.(..((((((....(((((.........----............)))))...))))......(((......))).))..).)))))...... ( -13.60) >DroSec_CAF1 2571 81 - 1 UAUGCUGGCCAUUAGGAUCUAUCACAAACAAAC----AUAUAUUA------------CUCAAAUAUAGCCAUAACCGGCAUAAAUUACAUAUUUUAU ((((((((((....)).................----.((((((.------------....)))))).......))))))))............... ( -13.00) >DroSim_CAF1 2403 93 - 1 UAUGCUGGCCAUUAGGAUAUAUCACAAACAAAC----AUAUAUUAUGGAUGAUACAACUCAAAUAUAGCCAUAACCGGCAUGAAACACAUAUUUCAU ((((((((((....)).................----.....((((((...(((.........)))..))))))))))))))............... ( -15.80) >DroEre_CAF1 2599 73 - 1 UACGCUGGCCAUUUGGAUAUAUCAUAAACAA-----CAUAUAUCA---------------AAAUAUAGCCAUAACCUACGUGAACU----AUUUCAC .....((((.((((.(((((((.........-----.))))))).---------------))))...))))........(((((..----..))))) ( -11.80) >DroYak_CAF1 2606 77 - 1 UAUGCUGGCGAUUAGCAUAUAUCAUAAAUAAACAUACAUGCAUCA----UGA-------CAAAUAUAGCCAUAACCGGCACUAACG---------AC (((((((.....)))))))..((.............((((...))----)).-------........(((......)))......)---------). ( -11.50) >consensus UAUGCUGGCCAUUAGGAUAUAUCACAAACAAAC____AUAUAUUA____UGA_____CUCAAAUAUAGCCAUAACCGGCAUGAACU_CAUAUUUCAU ((((((((......((.(((((........................................))))).))....))))))))............... ( -6.76 = -7.52 + 0.76)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:30 2006