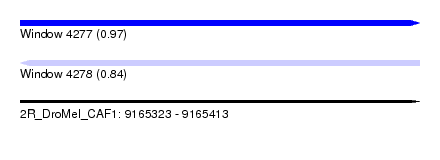

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,165,323 – 9,165,413 |
| Length | 90 |
| Max. P | 0.970243 |

| Location | 9,165,323 – 9,165,413 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 81.63 |
| Mean single sequence MFE | -29.23 |
| Consensus MFE | -24.52 |
| Energy contribution | -25.48 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.57 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.970243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
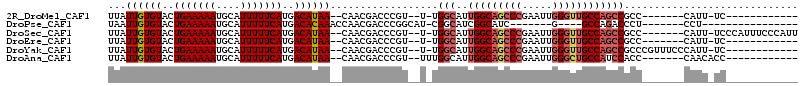
>2R_DroMel_CAF1 9165323 90 + 20766785 UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA--CAACGACCCGU--U-UGGCAUUGGCAGCCCGAAUUGGGUUGCCAGCCGCC-------CAUU-UC------------ ...((((((..(((((((....)))))))..))))))--.........(.--.-((((..(((((((((.....)))))))))))))..-------)...-..------------ ( -30.60) >DroPse_CAF1 34128 80 + 1 UAAUUGUGUACUGAAAAAUGCAUUUUUCAUGACACAAACCAACGACCCGGCAU-CGGCAUCGGCAUC-------G----GCCAGACCCU-------CCU---------------- ...((((((..(((((((....)))))))..))))))...........(((..-..((....))...-------.----))).......-------...---------------- ( -18.30) >DroSec_CAF1 49693 102 + 1 UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA--CAACGACCCGU--U-UGGCAUUGGCAGCCCGAAUUGGGUUGCCAGCCGCC-------CAUU-UCCCAUUUCCCAUU ...((((((..(((((((....)))))))..))))))--.........(.--.-((((..(((((((((.....)))))))))))))..-------)...-.............. ( -30.60) >DroEre_CAF1 42561 90 + 1 UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA--CAACGACCCGU--U-UGGCAUUGGCAGCCCGAAUUGGGUUGCCAGCCGCC-------CAUU-UC------------ ...((((((..(((((((....)))))))..))))))--.........(.--.-((((..(((((((((.....)))))))))))))..-------)...-..------------ ( -30.60) >DroYak_CAF1 40724 97 + 1 UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA--CAACGACCCGU--U-UGGCAUUGGCAGCCCGAAUUGGGUUGCCAGCCGCCCGUUUCCCAUU-UC------------ ...((((((..(((((((....)))))))..))))))--....((..((.--.-((((..(((((((((.....)))))))))))))..))..)).....-..------------ ( -33.90) >DroAna_CAF1 43948 92 + 1 UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA--CAACGACCCGU--UUUGGCAUUGGCAGCCCGAAUUGGGCUGCCAUCCACC-------CAACACC------------ ...((((((..(((((((....)))))))..))))))--.........((--.((((...(((((((((.....))))))))).....)-------))).)).------------ ( -31.40) >consensus UUAUUGUGUACUGAAAAAUGCAUUUUUCAUGACAUAA__CAACGACCCGU__U_UGGCAUUGGCAGCCCGAAUUGGGUUGCCAGCCGCC_______CAUU_UC____________ ...((((((..(((((((....)))))))..))))))..................(((..(((((((((.....))))))))))))............................. (-24.52 = -25.48 + 0.96)

| Location | 9,165,323 – 9,165,413 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 81.63 |
| Mean single sequence MFE | -28.62 |
| Consensus MFE | -19.87 |
| Energy contribution | -20.50 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.835279 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
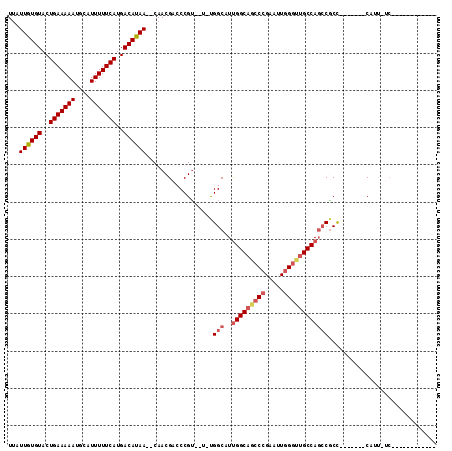
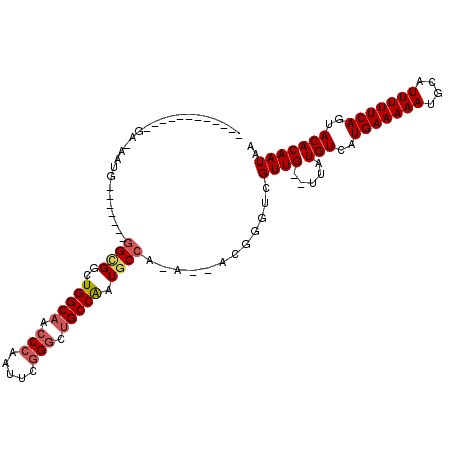
>2R_DroMel_CAF1 9165323 90 - 20766785 ------------GA-AAUG-------GGCGGCUGGCAACCCAAUUCGGGCUGCCAAUGCCA-A--ACGGGUCGUUG--UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ------------((-..((-------((((..(((((.(((.....))).))))).)))).-.--.))..)).(((--((.(((..(((((((....)))))))..))).))))) ( -27.40) >DroPse_CAF1 34128 80 - 1 ----------------AGG-------AGGGUCUGGC----C-------GAUGCCGAUGCCG-AUGCCGGGUCGUUGGUUUGUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUUA ----------------..(-------(.(..(((((----.-------...((....))..-..)))))..).))...((((((..(((((((....)))))))..))))))... ( -23.90) >DroSec_CAF1 49693 102 - 1 AAUGGGAAAUGGGA-AAUG-------GGCGGCUGGCAACCCAAUUCGGGCUGCCAAUGCCA-A--ACGGGUCGUUG--UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ..((.....((..(-((((-------((((..(((((.(((.....))).))))).)))).-.--.((.(.(((..--..))).).)).......)))))..)).....)).... ( -29.80) >DroEre_CAF1 42561 90 - 1 ------------GA-AAUG-------GGCGGCUGGCAACCCAAUUCGGGCUGCCAAUGCCA-A--ACGGGUCGUUG--UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ------------((-..((-------((((..(((((.(((.....))).))))).)))).-.--.))..)).(((--((.(((..(((((((....)))))))..))).))))) ( -27.40) >DroYak_CAF1 40724 97 - 1 ------------GA-AAUGGGAAACGGGCGGCUGGCAACCCAAUUCGGGCUGCCAAUGCCA-A--ACGGGUCGUUG--UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ------------((-..(((....).((((..(((((.(((.....))).))))).)))).-.--.))..)).(((--((.(((..(((((((....)))))))..))).))))) ( -31.00) >DroAna_CAF1 43948 92 - 1 ------------GGUGUUG-------GGUGGAUGGCAGCCCAAUUCGGGCUGCCAAUGCCAAA--ACGGGUCGUUG--UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ------------..((((.-------..(((.(((((((((.....)))))))))...))).)--))).....(((--((.(((..(((((((....)))))))..))).))))) ( -32.20) >consensus ____________GA_AAUG_______GGCGGCUGGCAACCCAAUUCGGGCUGCCAAUGCCA_A__ACGGGUCGUUG__UUAUGUCAUGAAAAAUGCAUUUUUCAGUACACAAUAA ..........................((((..(((((.(((.....))).))))).))))............((((.....(((..(((((((....)))))))..))))))).. (-19.87 = -20.50 + 0.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:03 2006