
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,131,741 – 9,131,851 |
| Length | 110 |
| Max. P | 0.780410 |

| Location | 9,131,741 – 9,131,851 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.48 |
| Mean single sequence MFE | -40.43 |
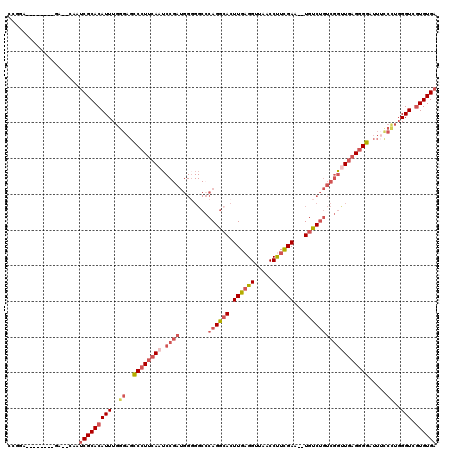
| Consensus MFE | -28.53 |
| Energy contribution | -30.70 |
| Covariance contribution | 2.17 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.780410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
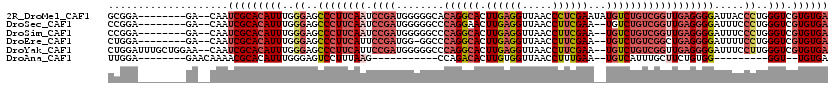
>2R_DroMel_CAF1 9131741 110 - 20766785 GCGGA--------GA--CAAUCGCACAUUUGGGAGCCCUUCAAUCCGAUGGGGGCACAGGCACUUGAGGUUAACCCUCGAAUAUGUCUGUCGGUUGAGGGGAUUACCCUGGGUCGUGUGA ..(..--------..--)..(((((((((..((..((((((((.((((((((.((....))..((((((.....)))))).....)))))))))))))))).....))..))).)))))) ( -45.60) >DroSec_CAF1 6929 108 - 1 CCGGA--------GA--CAAUCGCACAUUUGGGAGCCCUUCAAUCCGAUGGGGGCCCAGGAACUUGAGGUUAACCUUCGAA--UGUCUGUCGGUUGAGGGGAUUUCCCUGGGUCGUGUGA ..(..--------..--)..((((((((((((((.((((((((.(((((((....)).(((..((((((.....)))))).--..))))))))))))))))...)))).)))).)))))) ( -43.50) >DroSim_CAF1 6913 108 - 1 CCGGA--------GA--CAAUCGCACAUUUGGGAGCCCUUCAAUCCGAUGGGGGCCCAGGCACUUGAGGUUAACCUUCGAA--UGUCUGUCGGUUGAGGGGAUUUCCCUGGGUCGUGUGA ..(..--------..--)..((((((((((((((.((((((((.(((((((....))(((((.((((((.....)))))).--))))))))))))))))))...)))).)))).)))))) ( -48.10) >DroEre_CAF1 6741 107 - 1 CUGGA--------GA--CAAUCGCACAUUUGGGAGCCCUUCAUUCCGAUGG-GGCCCAGGCACUUGAGGUUAACCUUCGAA--UGUCUGUCGGCUGAGGGGAUUUUCCUGGGUCGUGUGA .((..--------..--)).(((((((((..(((.(((((((..((....)-)(((((((((.((((((.....)))))).--))))))..))))))))))....)))..))).)))))) ( -46.20) >DroYak_CAF1 6952 116 - 1 CUGGAUUUGCUGGAA--CAAUCGCACAUUUGGGAGCCCUUCAUUCCGAUGGGGGCCCAGGCACUUGAGGUUAACCUUCGAA--UGUCUGUCGGUUGAGGGGAUUUCCUUGGGUCGUGUGA (..(.....)..)..--...((((((((..((((.(((((((..(((((((....))(((((.((((((.....)))))).--)))))))))).)))))))...))).)..)).)))))) ( -44.00) >DroAna_CAF1 10044 88 - 1 UUGGA--------GAACAAAACGCACAUUUGGGAGUCCUUUAAG-----------CCAGACACUUGUGGUUAACCUUUGAA--UGUCAUUUGCUUCUGUGG---------GGU--UGUGA .....--------..((((..(.((((...((....))...(((-----------(..((((.(((.((....))..))).--))))....)))).)))))---------..)--))).. ( -15.20) >consensus CCGGA________GA__CAAUCGCACAUUUGGGAGCCCUUCAAUCCGAUGGGGGCCCAGGCACUUGAGGUUAACCUUCGAA__UGUCUGUCGGUUGAGGGGAUUUCCCUGGGUCGUGUGA ....................(((((((((..((..((((((((.((((........((((((.((((((.....))))))...)))))))))))))))))).....))..))).)))))) (-28.53 = -30.70 + 2.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:52 2006