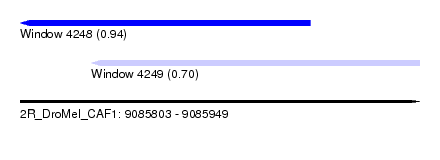

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 9,085,803 – 9,085,949 |
| Length | 146 |
| Max. P | 0.942293 |

| Location | 9,085,803 – 9,085,909 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 89.02 |
| Mean single sequence MFE | -27.88 |
| Consensus MFE | -23.94 |
| Energy contribution | -24.12 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.942293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
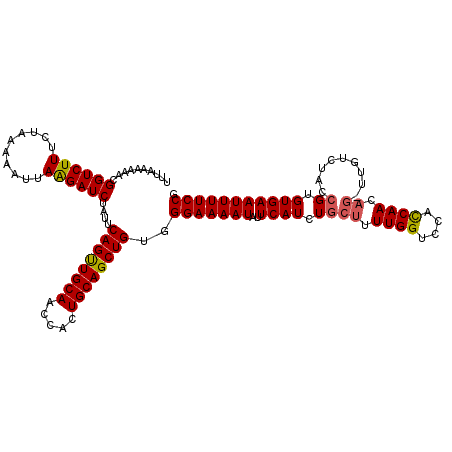
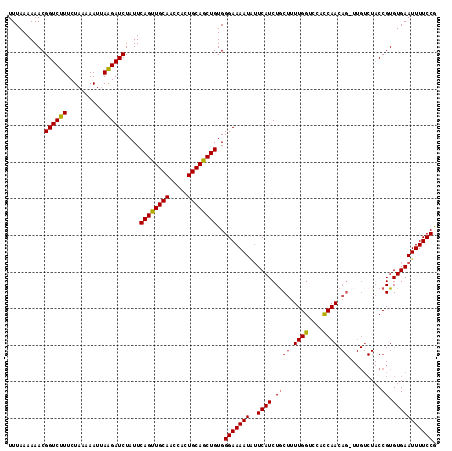
>2R_DroMel_CAF1 9085803 106 - 20766785 UUUAAAAAACGGUCUUUCUACAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUGUUCAUCUGCUUUUGGUCCACCAA-----------CCGUGUGAAUUUUCCG ..........((((((..........))))))....((((((((.....))))))))..(((((..((((((..((..((((....))))-----------..))))))))))))). ( -27.70) >DroSec_CAF1 50581 117 - 1 UUUAAAAAACGGUCUUUCUAAAAAUAAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUAUUCAUCUGCUUUUGGUCCUCCAACAGUUUGUCUACCGUGUGAAUUUUCCG ..........(((((((........)))))))....((((((((.....))))))))..(((((((..((((..(((.((((....)))).)))...........))))))))))). ( -27.52) >DroSim_CAF1 52794 117 - 1 UUUAAAAAACGGUCUUUCUAAAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUUUUCAUCUACUUUUGGUCCUUCAACAGAUUGUCUACCGUGUGAAUUUUCCG ..........((((((..........))))))....((((((((.....))))))))..(((((((((.(((.......(((((........)))))......))).))))))))). ( -25.82) >DroEre_CAF1 57902 116 - 1 UUUAAAAAACGGUCUUUCUAAAAAUUAGGAUCUAUUCAGCUGCAACCACUGCAGCUGUGGGAAAAUAGUCAUCUGCUUUUGGAGGACCAACAG-UUGUCUACCGUGUGAAUUUUCCA ..........((((((..........))))))....((((((((.....))))))))..(((((((..((((.((....((((.(((.....)-)).)))).)).))))))))))). ( -30.50) >consensus UUUAAAAAACGGUCUUUCUAAAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUAUUCAUCUGCUUUUGGUCCACCAACAG_UUGUCUACCGUGUGAAUUUUCCG ..........((((((..........))))))....((((((((.....))))))))..(((((((..((((.((((.((((....)))).)).........)).))))))))))). (-23.94 = -24.12 + 0.19)

| Location | 9,085,829 – 9,085,949 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.61 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -18.76 |
| Energy contribution | -18.70 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697060 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
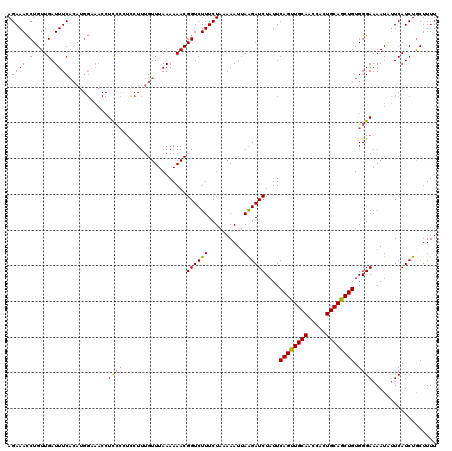
>2R_DroMel_CAF1 9085829 120 - 20766785 AGAAACCUGUUGAUUUCACAUGGAAACCUUACGUCCUUUGUUUAAAAAACGGUCUUUCUACAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUGUUCAUCUGCUUUU (((((.(((((..((..(((.(((.........)))..)))..))..)))))..)))))...........(((...((((((((.....))))))))..))).................. ( -23.30) >DroSec_CAF1 50618 120 - 1 AGAAACCUGUUGAUUUCACAUGGUAACCUCCCCUUCUUUGUUUAAAAAACGGUCUUUCUAAAAAUAAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUAUUCAUCUGCUUUU .(((((((((.(....)))).)))....((((..................(((((((........)))))))....((((((((.....)))))))).)))).....))).......... ( -25.80) >DroSim_CAF1 52831 120 - 1 AGAAACCUGUUGAUUUCACAUGGAAACCUCCCCGCCUUUGUUUAAAAAACGGUCUUUCUAAAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUUUUCAUCUACUUUU .((((.(....).))))..(((((((..((((..................((((((..........))))))....((((((((.....)))))))).))))...)))))))........ ( -25.20) >DroEre_CAF1 57938 109 - 1 AGAAAUCUGUUGAUUUC-----------UCCAUUUCUUUGUUUAAAAAACGGUCUUUCUAAAAAUUAGGAUCUAUUCAGCUGCAACCACUGCAGCUGUGGGAAAAUAGUCAUCUGCUUUU .((((((....))))))-----------(((...................((((((..........))))))....((((((((.....))))))))..))).................. ( -25.40) >consensus AGAAACCUGUUGAUUUCACAUGGAAACCUCCCCUCCUUUGUUUAAAAAACGGUCUUUCUAAAAAUUAAGAUCUAUUCAGUUGCAACCACUGCAGCUGUGGGAAAAUAUUCAUCUGCUUUU .((((.(....).))))...........((((..................((((((..........))))))....((((((((.....)))))))).)))).................. (-18.76 = -18.70 + -0.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:36 2006