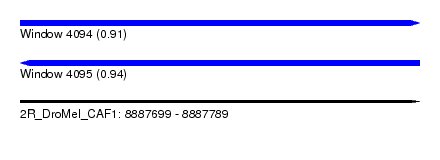
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,887,699 – 8,887,789 |
| Length | 90 |
| Max. P | 0.935775 |

| Location | 8,887,699 – 8,887,789 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 82.96 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -18.26 |
| Energy contribution | -18.32 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.13 |
| Mean z-score | -3.39 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.910077 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
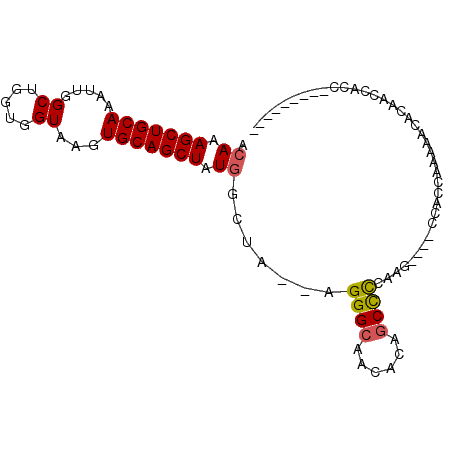
>2R_DroMel_CAF1 8887699 90 + 20766785 ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA--AGGGCAACACAGCCCCAAG----CCACCAAAAACACAACCACCCACAGCACU .....(((((......)).((((((..(((......(((((.--.((((......))))..))----))).......))).))))))...)))... ( -30.02) >DroSec_CAF1 132498 81 + 1 ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA--AGGGCAACACAGCCCCAAG----CCACCAAAAACACAACCACC--------- ....((((.......))))((((((..(((......(((((.--.((((......))))..))----))).......))).))))))--------- ( -28.22) >DroSim_CAF1 132370 81 + 1 ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA--AGGGCAACACAGCCCCAAG----CCACCAAAAACACAACCACC--------- ....((((.......))))((((((..(((......(((((.--.((((......))))..))----))).......))).))))))--------- ( -28.22) >DroEre_CAF1 135790 81 + 1 ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA--AGGGCAACACAGCCCCAAA----CCACCAAAAACACAACCACC--------- ....((((.......))))((((((......(((....))).--.((((......))))...)----)))))...............--------- ( -22.90) >DroYak_CAF1 137842 90 + 1 ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA--AGGGCAACACAGCCCCAAA----CCACCGAAAACACAACCACCCACAGCACU .....((((.....(((..((((((......(((....))).--.((((......))))...)----)))))(......)..)))....))))... ( -25.30) >DroAna_CAF1 131234 82 + 1 ACAAAGCUGCAAAUUGGCUGGCGGUAAGUGCAGCUAUAGCUUCUGGGGCAAUACUCCUUCAAUGGCCCCUUCGAUACCCCUU-------------- ....(((((((.....((.....))...))))))).........(((((...............))))).............-------------- ( -20.86) >consensus ACAAAGCUGCAAAUUGGCUGGUGGUAAGUGCAGCUAUGGCUA__AGGGCAACACAGCCCCAAG____CCACCAAAAACACAACCACC_________ .((.(((((((.....((.....))...))))))).)).......((((......))))..................................... (-18.26 = -18.32 + 0.06)

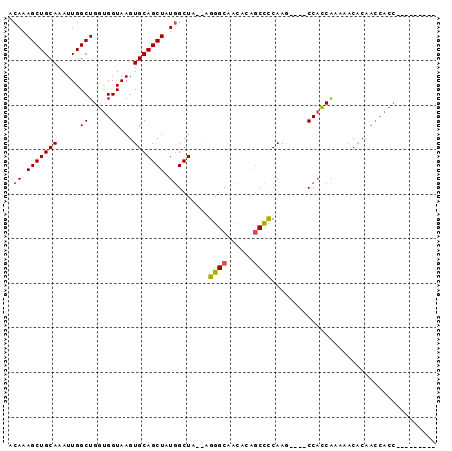
| Location | 8,887,699 – 8,887,789 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 82.96 |
| Mean single sequence MFE | -34.94 |
| Consensus MFE | -20.58 |
| Energy contribution | -22.17 |
| Covariance contribution | 1.58 |
| Combinations/Pair | 1.14 |
| Mean z-score | -4.07 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8887699 90 - 20766785 AGUGCUGUGGGUGGUUGUGUUUUUGGUGG----CUUGGGGCUGUGUUGCCCU--UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU ...(((((.((((((.((((.....((((----((.(((((......)))))--.))))))....))))..)))))).........)))))..... ( -40.90) >DroSec_CAF1 132498 81 - 1 ---------GGUGGUUGUGUUUUUGGUGG----CUUGGGGCUGUGUUGCCCU--UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU ---------((((((.((((.....((((----((.(((((......)))))--.))))))....))))..)))))).((......))........ ( -37.10) >DroSim_CAF1 132370 81 - 1 ---------GGUGGUUGUGUUUUUGGUGG----CUUGGGGCUGUGUUGCCCU--UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU ---------((((((.((((.....((((----((.(((((......)))))--.))))))....))))..)))))).((......))........ ( -37.10) >DroEre_CAF1 135790 81 - 1 ---------GGUGGUUGUGUUUUUGGUGG----UUUGGGGCUGUGUUGCCCU--UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU ---------((((((.((((.....((((----((.(((((......)))))--.))))))....))))..)))))).((......))........ ( -34.70) >DroYak_CAF1 137842 90 - 1 AGUGCUGUGGGUGGUUGUGUUUUCGGUGG----UUUGGGGCUGUGUUGCCCU--UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU ...(((((.((((((.((((.....((((----((.(((((......)))))--.))))))....))))..)))))).........)))))..... ( -38.50) >DroAna_CAF1 131234 82 - 1 --------------AAGGGGUAUCGAAGGGGCCAUUGAAGGAGUAUUGCCCCAGAAGCUAUAGCUGCACUUACCGCCAGCCAAUUUGCAGCUUUGU --------------..((((((((....)).((......)).....))))))....((...(((((((.................)))))))..)) ( -21.33) >consensus _________GGUGGUUGUGUUUUUGGUGG____CUUGGGGCUGUGUUGCCCU__UAGCCAUAGCUGCACUUACCACCAGCCAAUUUGCAGCUUUGU .........((((((.((((.....((((.......(((((......))))).....))))....))))..)))))).((......))........ (-20.58 = -22.17 + 1.58)

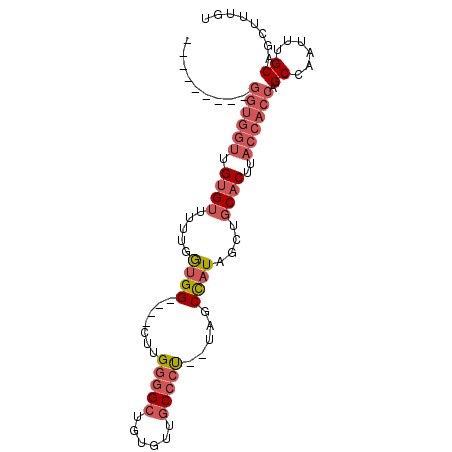
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:06 2006