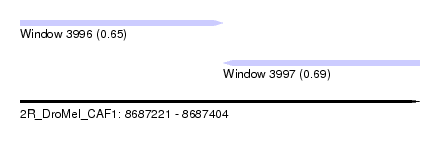
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,687,221 – 8,687,404 |
| Length | 183 |
| Max. P | 0.689088 |

| Location | 8,687,221 – 8,687,314 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 82.44 |
| Mean single sequence MFE | -31.14 |
| Consensus MFE | -20.05 |
| Energy contribution | -20.33 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.647458 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8687221 93 + 20766785 AGGCAAAGCUGCAAAAACAGCUGAU-----CGCAAGCGAAAGCUUUCCGUGCG------CAGCGGAGCUUUUGAGUUGGCCGGCA-GCUGAUAAAUGAGCACAGU---- ......((((((.....(((((.(.-----.((((((....))))(((((...------..)))))))...).)))))....)))-)))................---- ( -31.20) >DroSec_CAF1 2582 93 + 1 AGGCAAAGCUACAAAAACAGCUGAU-----CGCCAGCGAAAGCUUUCCGUGCG------CAGCGGAGCUUUUGAGUUGGCCGGCA-GCUGAUAAAUGAGCACAGU---- .......(((.......((((((..-----.((((((((((((..(((((...------..)))))))))))..))))))...))-)))).......))).....---- ( -35.54) >DroSim_CAF1 4979 93 + 1 AGGCAAAGCUGCAAAAACAGCUGAU-----CGCAAGCGAAAGCUUUCCGUGCG------CAACGGAGCUUUUGAGUUGGCCGGCA-GCUGAUAAAUGAGCACAGU---- ......((((((.....(((((.(.-----.((((((....))))(((((...------..)))))))...).)))))....)))-)))................---- ( -31.50) >DroEre_CAF1 2499 93 + 1 AGGCAAAGCUGCAAAAACAGCUGAU-----UGCUCGCGAAAGCUUUCCGUUCG------CCGCGGAGCUUUUGAGUUGGCCCGCA-GCUGAUAAAUGAGCACAGU---- ......(((((......)))))...-----(((((((....)).....(((.(------(.((((.(((........))))))).-)).)))....)))))....---- ( -32.10) >DroYak_CAF1 4924 93 + 1 AGGCAAAGCUACAAAAACAGCUGAU-----CGCAGGCGAAAGCUUUCCGUUCG------CCGCGCAGCUUUUGAGUUGGCCUGCA-GCUGAUAAAUGAGCACAGU---- ((((..((((.((((...(((((..-----(((.((((((.(.....).))))------)))))))))))))))))).))))...-(((........))).....---- ( -30.20) >DroPer_CAF1 3113 108 + 1 AAGAAAACUUGGGAUAACAGCUGUUUUGGACGCUA-CAAAAGCUCGUCGAGCGAGAAAGAAGCAAAGCUUUUGUGUGGGCUUGCAGGCUGAUAAAAAAGCACAGUACAG ....................((((.(((...(((.-(.....((((.....))))...).)))...((((((.(((.((((....)))).))).)))))).))).)))) ( -26.30) >consensus AGGCAAAGCUGCAAAAACAGCUGAU_____CGCAAGCGAAAGCUUUCCGUGCG______CAGCGGAGCUUUUGAGUUGGCCGGCA_GCUGAUAAAUGAGCACAGU____ .(((..(((((......)))))..............((((((((..((((...........)))))))))))).....))).....(((........)))......... (-20.05 = -20.33 + 0.28)

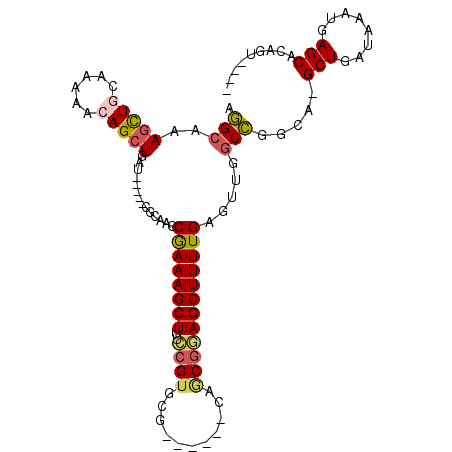
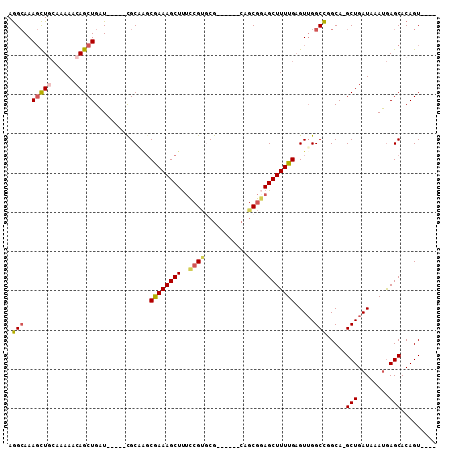
| Location | 8,687,314 – 8,687,404 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 80.85 |
| Mean single sequence MFE | -30.18 |
| Consensus MFE | -19.14 |
| Energy contribution | -19.38 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.689088 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8687314 90 - 20766785 AGAGGUGUCUCGGUUGA----ACGGUGGGAACUGAAGAAGGCGGCGUUGGCGAAGACGCUGCUGAUUUGUUUGCCUGUUGACCGACGAGCGACA ....((...((((((.(----((((.(.((((.(((...(((((((((......)))))))))..))))))).))))))))))))...)).... ( -34.20) >DroSec_CAF1 2675 86 - 1 AGAGGUGUCUCGGUUGA----ACGGGGGGAACUGAAGAAGGCGGCGGUGGCGAAGACGCUGCUGAU----UUGCCUGUUGACCGACGAGCGACA ....((...((((((.(----(((((.(((.((.....)).((((((((.......)))))))).)----)).))))))))))))...)).... ( -31.20) >DroSim_CAF1 5072 86 - 1 AGAGGUGUCUCGGUUGA----ACGGUGGGAACUGAAGAAGGCGGCGGUGGCGAAGACGCUGCUGGU----UUGCCUGUUGACCGACGAGCGACA ....((...(((((..(----.((((....))))....(((((((((..(((....)))..)))..----)))))).)..)))))...)).... ( -31.20) >DroEre_CAF1 2592 79 - 1 AG--------CGGUUGACAAGACAGGAGGAACUGAAGAAGGCGGCGGC---GAAGGCGCUGCUGGU----UUGCCUGUUGACCGACGAGCGACA .(--------(.((((....((((((..(((((......(((((((.(---....)))))))))))----)).))))))...))))..)).... ( -30.50) >DroYak_CAF1 5017 76 - 1 AG--------CGGUUGACAAUACAGGAGGAACUGAAAAAGGUGGCGUU------GGAGCUGCUGGU----UUGCCUGUUGACCGACGAGCGACA .(--------((((..(((...(((......)))(((..((..((...------...))..))..)----))...)))..)))).)........ ( -23.80) >consensus AGAGGUGUCUCGGUUGA____ACGGGGGGAACUGAAGAAGGCGGCGGUGGCGAAGACGCUGCUGGU____UUGCCUGUUGACCGACGAGCGACA ..........((((..(.....(((......)))....(((((((((((.......))))))).........)))).)..)))).......... (-19.14 = -19.38 + 0.24)

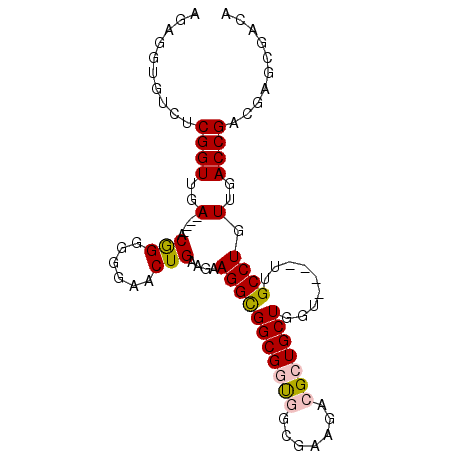
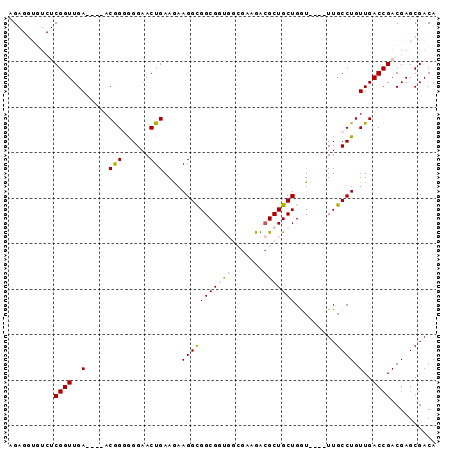
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:29:34 2006