
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,638,168 – 8,638,263 |
| Length | 95 |
| Max. P | 0.785121 |

| Location | 8,638,168 – 8,638,263 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.93 |
| Mean single sequence MFE | -27.95 |
| Consensus MFE | -16.73 |
| Energy contribution | -17.28 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.785121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8638168 95 - 20766785 ------------AAGAAUUUGGCAGCCA-CGA-GAGUCGCACGUCUCGCCAAUGGCUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUGGCCAGGCGGCAG-AAGG----------GGGU ------------....(((((.((((((-(((-((........)))))....)))))).)))))..((((((.((.(((...((.....))...))).)))-))))----------)... ( -33.60) >DroSec_CAF1 18301 75 - 1 ------------AAGAAUUUGGCAGCCA-CGA-GAAUCGCACGUCUCGCCAAUGGUUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUG------------------------------- ------------((((((((.((.((((-.((-(((......(((..((((.....))))....)))))))).)))))).)))))))).------------------------------- ( -21.50) >DroSim_CAF1 22017 75 - 1 ------------AAGAAUUUGGCAGCCA-CGA-GAAUCGCACGUCUCGCCAAUGGUUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUG------------------------------- ------------((((((((.((.((((-.((-(((......(((..((((.....))))....)))))))).)))))).)))))))).------------------------------- ( -21.50) >DroEre_CAF1 18168 75 - 1 ------------AAGAAUUUGGCAGCCA-CGA-GAGUCGCACGUCUCGCCAAUGGCUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUG------------------------------- ------------((((((((.((.((((-.((-((((((...(((..(((...))).)))...))))))))..)))))).)))))))).------------------------------- ( -24.70) >DroYak_CAF1 18326 100 - 1 ------------AAGAAUUUGGCAGCCA-CGA-GAGUCGCACGUCUCGCCAAUGGCUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUGGCCAGGCGGCAA-GGGGGCUU-----UGGGU ------------(((.(((((.((((((-(((-((........)))))....)))))).)))))..)))......((.(((((((((..(((....)))..-.)))))))-----)).)) ( -35.50) >DroAna_CAF1 18831 119 - 1 GAUUUCCCUUCGAAAUUUUUGGCAUCCAACGCCAAGCUGCACGUCUCGCCAAUGGCUGGCAAAUGACUUUUCAUUGCCCAAAACUUUUCAA-AGGAGAGAGAGAGAGAGUUGCAAAGGAU ......((((.......((((((.......)))))).((((..(((((((...))).(((((.(((....)))))))).....(((((...-..))))).....))))..)))))))).. ( -30.90) >consensus ____________AAGAAUUUGGCAGCCA_CGA_GAGUCGCACGUCUCGCCAAUGGCUGGCAAAUGACUUUUCAUGGCGCAAAGUUUUUG_______________________________ ............((((((((.((.((((.....((((((...(((..(((...))).)))...))))))....)))))).))))))))................................ (-16.73 = -17.28 + 0.56)

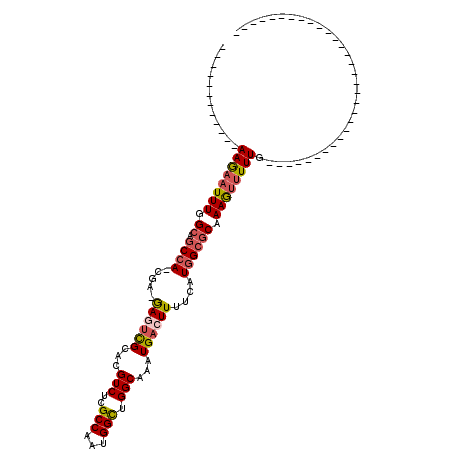
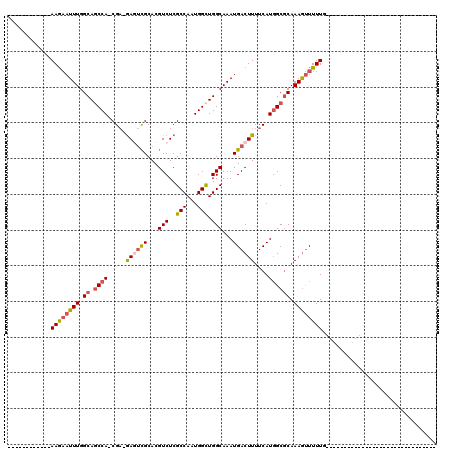
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:29:01 2006