
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,598,154 – 8,598,265 |
| Length | 111 |
| Max. P | 0.863379 |

| Location | 8,598,154 – 8,598,265 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 89.08 |
| Mean single sequence MFE | -41.42 |
| Consensus MFE | -29.93 |
| Energy contribution | -31.24 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8598154 111 + 20766785 CAGGAGCUG-----GAUAAAGGAGCAC-GGGGUGGAUUCCCUGCUCUGGACAGCGGACUGUGGA-UGUGGAUGUGGAUGUGGCAAAAGUUGCCAACUGGCCAACAAAUUGCCGCAUGA (((..((((-----......((((((.-((((.....))))))))))...))))...)))....-...........(((((((((..((((((....)).))))...))))))))).. ( -42.60) >DroSec_CAF1 82456 105 + 1 CAGGAGCUG-----GAUAAAGGAGCAC-GGGGUGGAUUCCGAGCUCUGGACAGCGGACUGUGGA-UG------UGGAUGUGGCAAAAGUAGCCAACUGGCCAACAAAUUGCCGCAUGA (((..((((-----......(((((.(-((((....))))).)))))...))))...)))....-..------...(((((((((..((.(((....)))..))...))))))))).. ( -38.50) >DroSim_CAF1 74418 111 + 1 CAGGAGCUG-----GAUAAAGGAGCAC-GGGGUGGAUUCCCUGCUCUGGACAGCGGACUGUGGA-UGUGAAUGUGGAUGUGGCAAAAGUUGCCAACUGGCCAACAAAUUGCCGCAUGA (((..((((-----......((((((.-((((.....))))))))))...))))...)))....-...........(((((((((..((((((....)).))))...))))))))).. ( -42.60) >DroYak_CAF1 77488 118 + 1 CAGGAGUUGGCCAGGAUAAAGGAGCUCUGGGGUGGAUUCCCUGCCCUGGACAGCGGACAACAGACUGUGCAUGUGGAUGAGGCAAAAGUUGCCAACUGGCCAACAAAUUGCCGCAUGA .....(((((((((.........((((..(((..(.....)..)))..))..))((.((((....(((.(((....)))..)))...))))))..))))))))).............. ( -42.00) >consensus CAGGAGCUG_____GAUAAAGGAGCAC_GGGGUGGAUUCCCUGCUCUGGACAGCGGACUGUGGA_UGUG_AUGUGGAUGUGGCAAAAGUUGCCAACUGGCCAACAAAUUGCCGCAUGA (((..((((...........(((((...((((.....)))).)))))...))))...)))................(((((((((..((((((....)).))))...))))))))).. (-29.93 = -31.24 + 1.31)


| Location | 8,598,154 – 8,598,265 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 89.08 |
| Mean single sequence MFE | -32.77 |
| Consensus MFE | -23.11 |
| Energy contribution | -23.30 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.747008 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8598154 111 - 20766785 UCAUGCGGCAAUUUGUUGGCCAGUUGGCAACUUUUGCCACAUCCACAUCCACA-UCCACAGUCCGCUGUCCAGAGCAGGGAAUCCACCCC-GUGCUCCUUUAUC-----CAGCUCCUG ..(((.(((((...((((.((....))))))..))))).)))...........-....(((...((((....((((((((.......)))-.))))).......-----))))..))) ( -32.30) >DroSec_CAF1 82456 105 - 1 UCAUGCGGCAAUUUGUUGGCCAGUUGGCUACUUUUGCCACAUCCA------CA-UCCACAGUCCGCUGUCCAGAGCUCGGAAUCCACCCC-GUGCUCCUUUAUC-----CAGCUCCUG ..(((.(((((...((.((((....))))))..))))).)))...------..-....(((...((((....((((.(((........))-).)))).......-----))))..))) ( -31.70) >DroSim_CAF1 74418 111 - 1 UCAUGCGGCAAUUUGUUGGCCAGUUGGCAACUUUUGCCACAUCCACAUUCACA-UCCACAGUCCGCUGUCCAGAGCAGGGAAUCCACCCC-GUGCUCCUUUAUC-----CAGCUCCUG ..(((.(((((...((((.((....))))))..))))).)))...........-....(((...((((....((((((((.......)))-.))))).......-----))))..))) ( -32.30) >DroYak_CAF1 77488 118 - 1 UCAUGCGGCAAUUUGUUGGCCAGUUGGCAACUUUUGCCUCAUCCACAUGCACAGUCUGUUGUCCGCUGUCCAGGGCAGGGAAUCCACCCCAGAGCUCCUUUAUCCUGGCCAACUCCUG ..............(((((((((..((((.....))))................((((.(((((........)))))(((......)))))))...........)))))))))..... ( -34.80) >consensus UCAUGCGGCAAUUUGUUGGCCAGUUGGCAACUUUUGCCACAUCCACAU_CACA_UCCACAGUCCGCUGUCCAGAGCAGGGAAUCCACCCC_GUGCUCCUUUAUC_____CAGCUCCUG ..(((.(((((...((((.((....))))))..))))).)))...............((((....))))...((((.(((.......)))...))))..................... (-23.11 = -23.30 + 0.19)
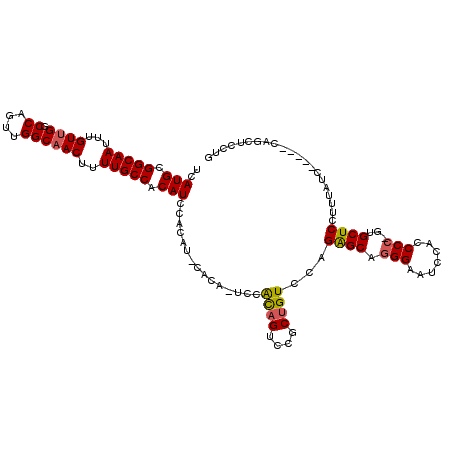



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:28:11 2006