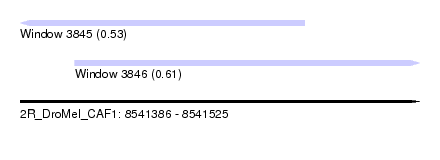

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,541,386 – 8,541,525 |
| Length | 139 |
| Max. P | 0.611450 |

| Location | 8,541,386 – 8,541,485 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 82.14 |
| Mean single sequence MFE | -22.70 |
| Consensus MFE | -16.06 |
| Energy contribution | -17.12 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.71 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.533099 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8541386 99 - 20766785 UAUCAGGCGGCAAUCAAUAAAACGCAAUAAUGCGAAUUGCAUUUGUGGAGCGGAGCGUUCCGAAUGCCCCAUAAAACUGAAACGGUUGGCAAGAGAAA----A ..((.((.((((.((........((((..((((.....))))))))((((((...)))))))).))))))....(((((...))))).....))....----. ( -24.80) >DroSec_CAF1 17021 103 - 1 UAUCAGGCGGCAAUCAAUAAAACGCAAUAAUGCGAAUUGCAUUUGUGGAGCGGAGCGGGCCGAACGCCCCAUAAAACUGAAACGGUUGGCAAGAAAAAAAAAA ........(.(((((.......((((....)))).......(((((((.(((...((...))..))).)))))))........))))).)............. ( -23.90) >DroSim_CAF1 20593 103 - 1 UAUCAGGCGGCAAUCAAUAAAACGCAAUAAUGCGAAUUGCAUUUGUGGAGCGGAGCGGGCCAAACGCCCCAUAAAACUGAAACGGUUGGCAAAAAGAAAAAAA ........(.(((((.......((((....)))).......(((((((.(((............))).)))))))........))))).)............. ( -23.00) >DroEre_CAF1 6476 78 - 1 UAUCAGGCGGCAAUCAAUAAUACGCAAUAAUGCGAAUUGCAUUUGUGGAGCGGAGCGGGCCGAACGGUCCAUAAA-------------------------AAA ..((...(.((((....((((.((((....)))).))))...)))).)....))..(((((....))))).....-------------------------... ( -19.10) >consensus UAUCAGGCGGCAAUCAAUAAAACGCAAUAAUGCGAAUUGCAUUUGUGGAGCGGAGCGGGCCGAACGCCCCAUAAAACUGAAACGGUUGGCAAGAAAAA__AAA ..((((...(((((........((((....)))).))))).(((((((.(((............))).))))))).))))....................... (-16.06 = -17.12 + 1.06)


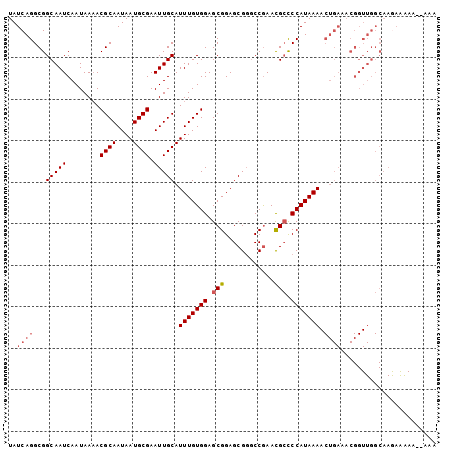
| Location | 8,541,405 – 8,541,525 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.36 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -22.14 |
| Energy contribution | -22.50 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.611450 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
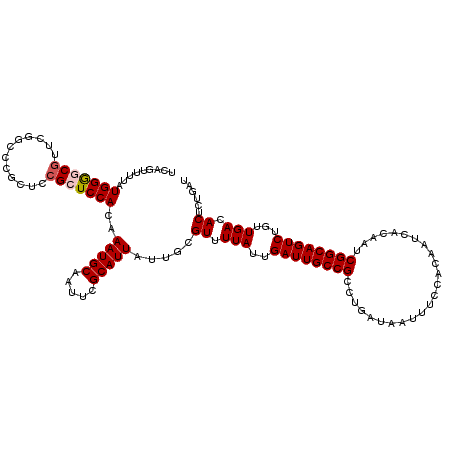
>2R_DroMel_CAF1 8541405 120 + 20766785 UCAGUUUUAUGGGGCAUUCGGAACGCUCCGCUCCACAAAUGCAAUUCGCAUUAUUGCGUUUUAUUGAUUGCCGCCUGAUAAUUUCCACAAUCACAAUCGGCAGUCUGUUGACACUCUGAU ((((.....((((((....(((....))))))))).(((((((((.......)))))))))....((((((((..((((..........))))....))))))))..........)))). ( -31.70) >DroSec_CAF1 17044 120 + 1 UCAGUUUUAUGGGGCGUUCGGCCCGCUCCGCUCCACAAAUGCAAUUCGCAUUAUUGCGUUUUAUUGAUUGCCGCCUGAUAUUUUCCACAAUCACAAUCGGCAGUCUGUUGACACUCUGAU ((((......((((((.......))))))(.((.(((((((((((.......)))))))))....((((((((..((((..........))))....)))))))).)).)))...)))). ( -32.90) >DroSim_CAF1 20616 120 + 1 UCAGUUUUAUGGGGCGUUUGGCCCGCUCCGCUCCACAAAUGCAAUUCGCAUUAUUGCGUUUUAUUGAUUGCCGCCUGAUAAUUUCCACAAUCACAAUCGGCAGUCUGUUGACACUCUGAU ((((......((((((.......))))))(.((.(((((((((((.......)))))))))....((((((((..((((..........))))....)))))))).)).)))...)))). ( -32.90) >DroEre_CAF1 6479 109 + 1 -----UUUAUGGACCGUUCGGCCCGCUCCGCUCCACAAAUGCAAUUCGCAUUAUUGCGUAUUAUUGAUUGCCGCCUGAUAAUUUCCAC------AAUCGGCAGUCUGUUGACACUCUGAU -----.....((.((....)).)).....(.((.(((.(((((((.......)))))))......((((((((..((..........)------)..))))))))))).)))........ ( -23.50) >DroYak_CAF1 20246 105 + 1 UCAGUUUUAUGGACCGUU---------UCGCUCCACAAAUGCAAUUCGCAUUAUUGGGUAUUAUUGAUUGCCGACUGAUAAUUUCCAC------AAUCGGCAGUCUGUUGACACUCUGAU .........((((.((..---------.)).))))..(((((.....)))))...((((.(((..(((((((((.((..........)------).)))))))))...))).)))).... ( -26.60) >consensus UCAGUUUUAUGGGGCGUUCGGCCCGCUCCGCUCCACAAAUGCAAUUCGCAUUAUUGCGUUUUAUUGAUUGCCGCCUGAUAAUUUCCACAAUCACAAUCGGCAGUCUGUUGACACUCUGAU .........(((((((............)))))))..(((((.....))))).....((.(((..((((((((........................))))))))...))).))...... (-22.14 = -22.50 + 0.36)

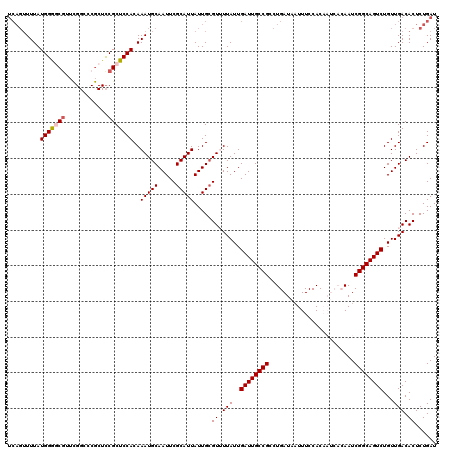
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:27:09 2006