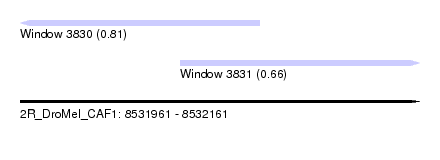
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,531,961 – 8,532,161 |
| Length | 200 |
| Max. P | 0.809784 |

| Location | 8,531,961 – 8,532,081 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.40 |
| Mean single sequence MFE | -25.45 |
| Consensus MFE | -23.38 |
| Energy contribution | -23.44 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.809784 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
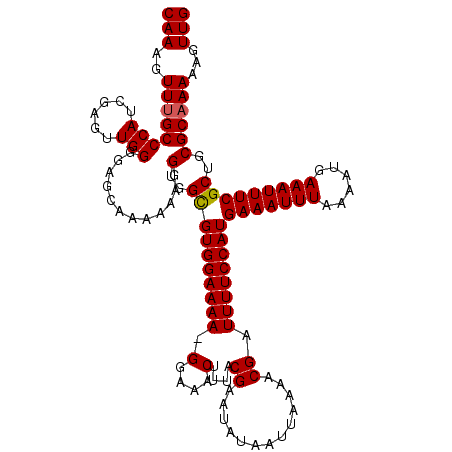
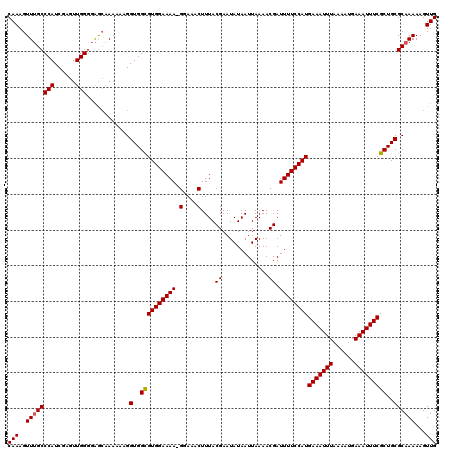
>2R_DroMel_CAF1 8531961 120 - 20766785 CAAAGUUUGCCCAUCGAGUUGGUGAGCAAAAAAGGUGGCGUGGAAAAAGAAAACUUUACGAAUAUAAUUAAAACGAUUUUCCAUGAAAUUUAAAAUGAAAUUUCGCUGCGCAAAAAGUUG .....(((((.((((.....)))).)))))....(..((((((((((.............................))))))))(((((((......)))))))))..)........... ( -23.95) >DroSec_CAF1 7408 119 - 1 CAAAGUUUGCCCAUCGAGUUGGGGAGCAAAAAAGGUGGCGUGGAAAA-GGAAACUUUACGAAUAUAAUUAAAACGAUUUUCCAUGAAAUUUAAAAUGAAAUUUCGCUGCGCAAAAAGUUG ....((((.((((......)))))))).......(..((((((((((-(....)))).((.............))....)))))(((((((......)))))))))..)........... ( -27.42) >DroSim_CAF1 10749 119 - 1 CAAAGUUUGCCCAUCGAGUUGGGGAGCGAAAAAGGUGGUGUGGAAAA-GGAAACUUUACGAAUAUAAUUAAAACGAUUUUCCAUGAAAUUUAAAAUGAAAUUUCGCUGCGCAAAAAGUUG (((..((((((((......)))(.(((............((((((((-(....)))).((.............))....)))))(((((((......)))))))))).))))))...))) ( -25.62) >DroYak_CAF1 10536 119 - 1 CAAAGUUAGCCCAUCAAGUUGGGAGAGAAAAAAGGUGGCGUGGAAAA-GGAAACUUUACGAAUAUAAUUAAAACGAUUUUCCAUGAAAUUUAAAAUGAAAUUUCGCUGCGCAAAAAGUUG .....((..((((......))))..)).......(..((((((((((-(....)))).((.............))....)))))(((((((......)))))))))..)........... ( -24.82) >consensus CAAAGUUUGCCCAUCGAGUUGGGGAGCAAAAAAGGUGGCGUGGAAAA_GGAAACUUUACGAAUAUAAUUAAAACGAUUUUCCAUGAAAUUUAAAAUGAAAUUUCGCUGCGCAAAAAGUUG (((..((((((((......)))............(..((((((((((.(....)....((.............)).))))))))(((((((......)))))))))..))))))...))) (-23.38 = -23.44 + 0.06)

| Location | 8,532,041 – 8,532,161 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.25 |
| Mean single sequence MFE | -26.48 |
| Consensus MFE | -23.55 |
| Energy contribution | -23.55 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.658288 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
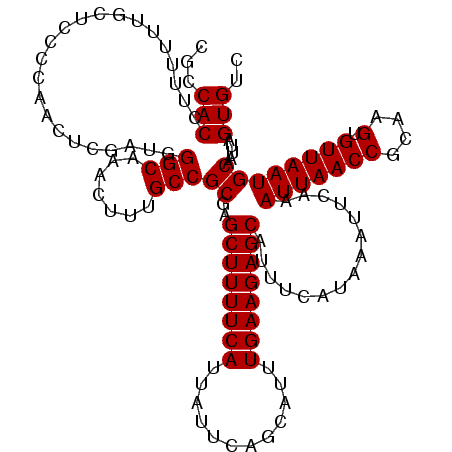
>2R_DroMel_CAF1 8532041 120 + 20766785 CGCCACCUUUUUUGCUCACCAACUCGAUGGGCAAACUUUGCCGCGAGCUUUUCAUUAUUCAGCAUUUGAAGAGCAUUUCAUAAAUUCAAAUUAACCGCAAGUGUUAAUGCAAUUAGUGUC (((.......((((((((.(.....).)))))))).......))).((((((((............))))))))....(((.((((...(((((((....).))))))..)))).))).. ( -28.54) >DroSec_CAF1 7487 120 + 1 CGCCACCUUUUUUGCUCCCCAACUCGAUGGGCAAACUUUGCCGCGAGCUUUUCAUUAUUCAGCAUUUGAAGAGCAUUUCAUAAAUUCAAAUUAACCGCAAGUGUUAAUGCAAUUAGUGUC ...(((.....((((........(((...((((.....)))).)))((((((((............))))))))...............(((((((....).))))))))))...))).. ( -26.70) >DroSim_CAF1 10828 120 + 1 CACCACCUUUUUCGCUCCCCAACUCGAUGGGCAAACUUUGCCGCGAGCUUUUCAUUAUUCAGCAUUUGAAGAGCAUUUCAUAAAUUCAAAUUAACCGCAAGUGUUAAUGCAAUUAGUGUC ...........((((....((......))((((.....))))))))((((((((............))))))))....(((.((((...(((((((....).))))))..)))).))).. ( -26.80) >DroYak_CAF1 10615 120 + 1 CGCCACCUUUUUUCUCUCCCAACUUGAUGGGCUAACUUUGCCGCGAGCUUUUCAUUAUUCAGCAUUUGAAGAGCAUUUCAUAAAUUCAAAUUAACCGCAAGUGUUAAUGCAAUUAGUGUC (((................((......))(((.......)))))).((((((((............))))))))....(((.((((...(((((((....).))))))..)))).))).. ( -23.90) >consensus CGCCACCUUUUUUGCUCCCCAACUCGAUGGGCAAACUUUGCCGCGAGCUUUUCAUUAUUCAGCAUUUGAAGAGCAUUUCAUAAAUUCAAAUUAACCGCAAGUGUUAAUGCAAUUAGUGUC ...(((.......................(((.......)))((..((((((((............))))))))...............(((((((....).)))))))).....))).. (-23.55 = -23.55 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:26:55 2006