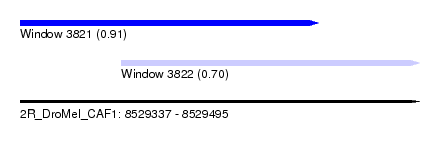
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,529,337 – 8,529,495 |
| Length | 158 |
| Max. P | 0.909108 |

| Location | 8,529,337 – 8,529,455 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.22 |
| Mean single sequence MFE | -24.00 |
| Consensus MFE | -23.40 |
| Energy contribution | -23.65 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.909108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
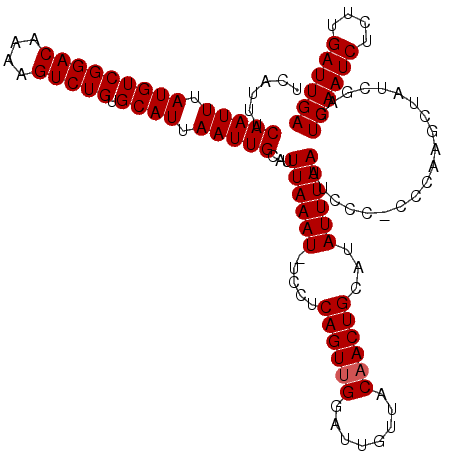
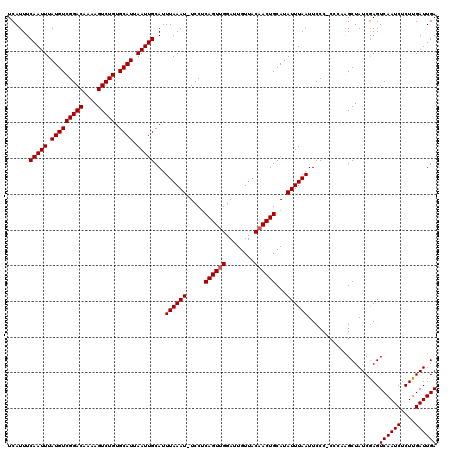
>2R_DroMel_CAF1 8529337 118 + 20766785 UCAUUUCAAUUUAUGUCGGACAAAAGUCUGUGCAUUAAUUGCAUUUAAAU-UCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCCCAGCUAUCGAGUCAAUCUCUUGAUUGA ((....(((((.(((((((((....))))).)))).)))))...((((((-....((((((........))))))...)))))).....-...........)).((((((....)))))) ( -25.10) >DroSec_CAF1 4869 119 + 1 UCAUUUCAAUUUAUGUCGGACAAAAGUCUGUGCAUUAAUUGCAUUUAAAUUUCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCCAGGCUAUCGAGUCAAUCUCUUGAUUGA ...............((((.....((((((((((.....)))).((((((.....((((((........))))))...)))))).....-..)))))).)))).((((((....)))))) ( -24.70) >DroSim_CAF1 8118 119 + 1 UCAUUUCAAUUUAUGUCGGACAAAAGUCUGUGCAUUAAUUGCAUUUAAAU-UCCUCAGUCGGAUUGUUACAACUGCAUAUUUAAUUCCCCCCCAGGCUAUCGAGUCAAUCUCUUGAUUGA ...............((((.....((((((((((.....)))).((((((-....((((.(........).))))...))))))........)))))).)))).((((((....)))))) ( -21.20) >DroYak_CAF1 7829 118 + 1 UCAUUUCAAUUUAUGUCGGACAAAAGUCUGUGCAUUAAUUGCAUUUAAAU-UCCCCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCAUAGCUAUCAAGUCAAUCUCUUGAUUGA ......(((((.(((((((((....))))).)))).)))))...((((((-....((((((........))))))...)))))).....-..............((((((....)))))) ( -25.00) >consensus UCAUUUCAAUUUAUGUCGGACAAAAGUCUGUGCAUUAAUUGCAUUUAAAU_UCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC_CCCAAGCUAUCGAGUCAAUCUCUUGAUUGA ......(((((.(((((((((....))))).)))).)))))...((((((.....((((((........))))))...))))))....................((((((....)))))) (-23.40 = -23.65 + 0.25)

| Location | 8,529,377 – 8,529,495 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.96 |
| Mean single sequence MFE | -24.27 |
| Consensus MFE | -20.89 |
| Energy contribution | -21.45 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.699747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
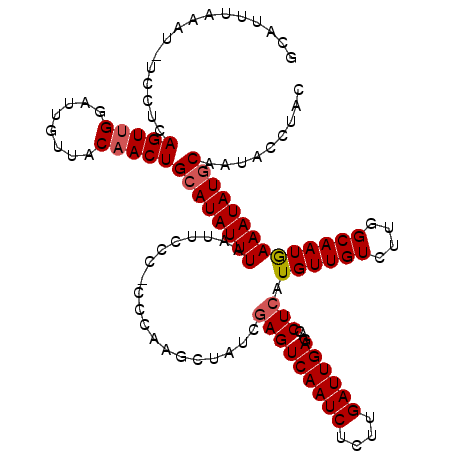
>2R_DroMel_CAF1 8529377 118 + 20766785 GCAUUUAAAU-UCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCCCAGCUAUCGAGUCAAUCUCUUGAUUGAGACCUCAUGUUGUCUUGGCAAUAAAAUAUGCAAUACCUAC ..........-.....(((((........)))))((((((((.(((((.-...((((.((.(((((((((....))))))...)))))))))....)).))).))))))))......... ( -26.00) >DroSec_CAF1 4909 119 + 1 GCAUUUAAAUUUCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCCAGGCUAUCGAGUCAAUCUCUUGAUUGAGACCUCAUGUUGUCUUGGCAAUGAAAUAUACAAUACCUAC ....((((((.....((((((........))))))...)))))).....-...(((.....(((((((((....))))))...)))..(((((....))))).............))).. ( -22.00) >DroSim_CAF1 8158 119 + 1 GCAUUUAAAU-UCCUCAGUCGGAUUGUUACAACUGCAUAUUUAAUUCCCCCCCAGGCUAUCGAGUCAAUCUCUUGAUUGAGACCUCAUGUUGUCUUGGCAAUGAAAUAUGCAAUAUCUAC .....(((..-(((......)))...)))....(((((((((.(((.((....((((....(((((((((....))))))...))).....)))).)).))).)))))))))........ ( -24.00) >DroYak_CAF1 7869 118 + 1 GCAUUUAAAU-UCCCCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC-CCAUAGCUAUCAAGUCAAUCUCUUGAUUGAGACCUCAUGUUGUCUUGGCAAUGAAAUAUGCAAUACCUAC ((((((((((-....((((((........))))))...)))))).....-.(((.((((.....((((((....))))))(((........))).)))).)))....))))......... ( -25.10) >consensus GCAUUUAAAU_UCCUCAGUUGGAUUGUUACAACUGCAUAUUUAAUUCCC_CCCAAGCUAUCGAGUCAAUCUCUUGAUUGAGACCUCAUGUUGUCUUGGCAAUGAAAUAUGCAAUACCUAC ................(((((........)))))((((((((...................(((((((((....))))))...))).((((((....))))))))))))))......... (-20.89 = -21.45 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:26:47 2006