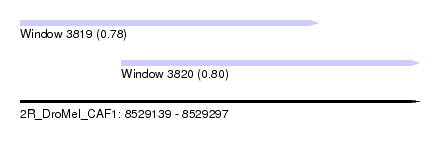
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,529,139 – 8,529,297 |
| Length | 158 |
| Max. P | 0.795076 |

| Location | 8,529,139 – 8,529,257 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.29 |
| Mean single sequence MFE | -42.08 |
| Consensus MFE | -34.35 |
| Energy contribution | -34.85 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8529139 118 + 20766785 CGGUAGCUUCGCGGUGCGUGAGAAAAGAUACACUCGGUAUCAGUUCGAGAUACUUAUG--UAUCUGCAGCUGUAUUUGUAGCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGC ..((.(((((((((((((((((....(((((.....))))).(((...))).))))))--))))((((((((......))))))))))))(((((((((...))))))))).))).)).. ( -45.30) >DroSec_CAF1 4669 120 + 1 CGGUAGCUUUGCGGUGCGUGAGAAAAGAUACACUCGACAUCAGUUCGAGAUACUUAUGUGUAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCUC .(((.(((((((.((.(....)...(((((((........(((((..((((((......))))))..)))))....))))))))).))))(((((((((...))))))))).))).))). ( -39.60) >DroSim_CAF1 7918 120 + 1 CGGUAGCUUUGCGGUGCGUGAGAAAAGAUACACUCGACAGCAGUUCGAGAUACUUAUGUGUAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCUC .(((.(((((((.((.(....)...(((((((....(((((.((...((((((......)))))))).)))))...))))))))).))))(((((((((...))))))))).))).))). ( -43.70) >DroYak_CAF1 7631 118 + 1 CGGUAGCUUCGCGGUGCGUGAGAGAAGAUACACUCGGCAUCAGUUCAAGAUACUUAUG--UAUCUGUAGCUGUAUUUGUAUCUGCAGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGC ..((.(((((((.((.(....)...(((((((........(((((..((((((....)--)))))..)))))....))))))))).))))(((((((((...))))))))).))).)).. ( -39.70) >consensus CGGUAGCUUCGCGGUGCGUGAGAAAAGAUACACUCGACAUCAGUUCGAGAUACUUAUG__UAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGC .....(((..((((((((((((..........))))....(((((..((((((......))))))..)))))....)))).)))).((..(((((((((...)))))))))..))))).. (-34.35 = -34.85 + 0.50)

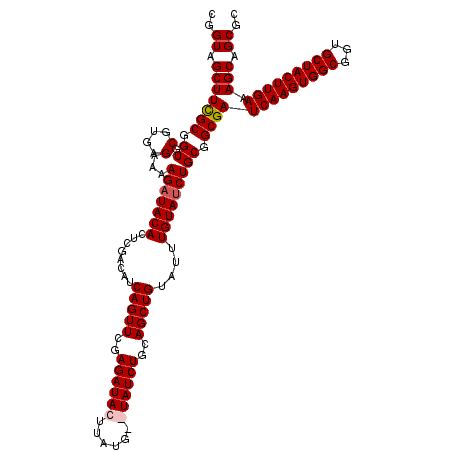
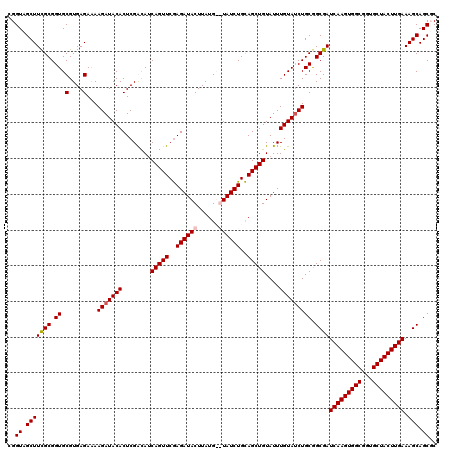
| Location | 8,529,179 – 8,529,297 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.43 |
| Mean single sequence MFE | -34.47 |
| Consensus MFE | -28.35 |
| Energy contribution | -29.35 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.795076 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8529179 118 + 20766785 CAGUUCGAGAUACUUAUG--UAUCUGCAGCUGUAUUUGUAGCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGCUGCGAGUUCCUCAAAUUCACUUCUAAUUGAUCCAAAAUUG ..(((..((((((....)--)))))(((((((......))))))))))(((((((..(((.((((.......)))).)))..)(((((.....)))))........))))))........ ( -40.30) >DroSec_CAF1 4709 120 + 1 CAGUUCGAGAUACUUAUGUGUAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCUCACCGAGUUCCUCAAAUUCACUUCUAAUUGAUUCAAAAUUG ......(((..((((..(((...((((.((((((........))))))..(((((((((...)))))))))..))))..))).))))..)))............................ ( -33.80) >DroSim_CAF1 7958 120 + 1 CAGUUCGAGAUACUUAUGUGUAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCUCACCGAGUUCCUCAAAUUCACUUCUAAUUGAUUCAAAAUUG ......(((..((((..(((...((((.((((((........))))))..(((((((((...)))))))))..))))..))).))))..)))............................ ( -33.80) >DroYak_CAF1 7671 118 + 1 CAGUUCAAGAUACUUAUG--UAUCUGUAGCUGUAUUUGUAUCUGCAGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGCUCCAAGUUCCUAAAAUUCACUUCUAAUUGAUUCAAAAUUG (((((..((((((....)--)))))..)))))..((((.(((.((.((..(((((((((...)))))))))..)).)).....((((...........))))......))).)))).... ( -30.00) >consensus CAGUUCGAGAUACUUAUG__UAUCUGCAGCUGUAUUUGUAUCUGCGGCGAUCAAGUGGCGGUGCUACUUGAAAGCAGCGCACCGAGUUCCUCAAAUUCACUUCUAAUUGAUUCAAAAUUG (((((..((((((......))))))..)))))..((((.(((.((.((..(((((((((...)))))))))..)).)).....(((((.....)))))..........))).)))).... (-28.35 = -29.35 + 1.00)

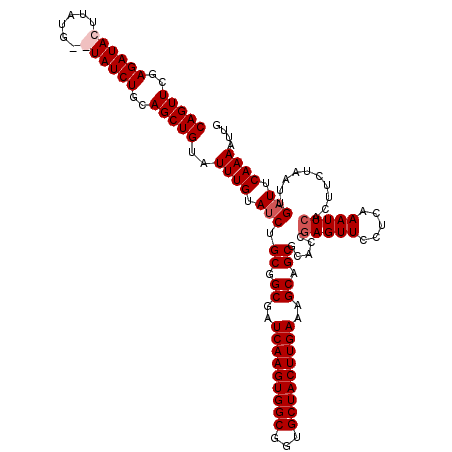
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:26:45 2006