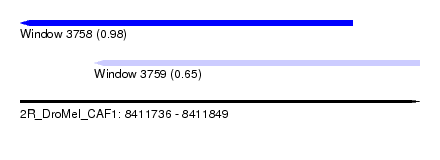

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,411,736 – 8,411,849 |
| Length | 113 |
| Max. P | 0.980330 |

| Location | 8,411,736 – 8,411,830 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 88.95 |
| Mean single sequence MFE | -25.45 |
| Consensus MFE | -21.92 |
| Energy contribution | -22.28 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980330 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
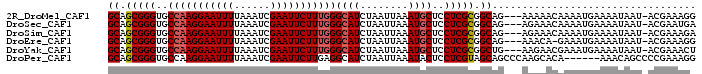
>2R_DroMel_CAF1 8411736 94 - 20766785 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCAG---AAAAACAAAAUGAAAAUAAU-ACGAAAGG ((.(((((..(((((((((((......)))))))))))((((........))))..))))).))..---....................-.(....). ( -26.00) >DroSec_CAF1 23246 94 - 1 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCAG---AGAAACAAAAUGAAAAUAAU-ACGAAUGA ((.(((((..(((((((((((......)))))))))))((((........))))..))))).))..---....................-........ ( -24.60) >DroSim_CAF1 22882 94 - 1 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCAG---AGAAACAAAAUGAAAAUAAU-ACGAAAGA ((.(((((..(((((((((((......)))))))))))((((........))))..))))).))..---....................-.(....). ( -25.80) >DroEre_CAF1 25129 93 - 1 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCAG---AAACA-GAAAUGAAAAUAAU-ACGAAAGG ((.(((((..(((((((((((......)))))))))))((((........))))..))))).))..---...((-....))........-.(....). ( -27.20) >DroYak_CAF1 23186 94 - 1 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCUG---AAGAACGAAAUGAAAAUAAU-ACGAAACU .((((((((((((((((((((......))))))))).))))))).......(((.....)))))))---....................-........ ( -24.70) >DroPer_CAF1 22573 92 - 1 GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUGAGGCAUCUAAUUAAAUACUCCUCGUAGCAGCCCAAGCACA------AAACAGCCCCGAAAGG ((.(((((((((.((((((((......))))))))..))))))).......(((.....))))).((....))...------.....)).((....)) ( -24.40) >consensus GCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUGGGCAUCUAAUUAAAUGCUCCUCGCGGCAG___AAAAACAAAAUGAAAAUAAU_ACGAAAGG ((.(((((..(((((((((((......)))))))))))((((........))))..))))).)).................................. (-21.92 = -22.28 + 0.36)


| Location | 8,411,757 – 8,411,849 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 79.21 |
| Mean single sequence MFE | -27.07 |
| Consensus MFE | -14.51 |
| Energy contribution | -15.03 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652606 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
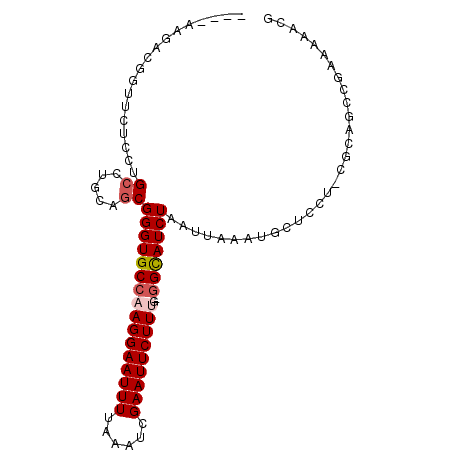
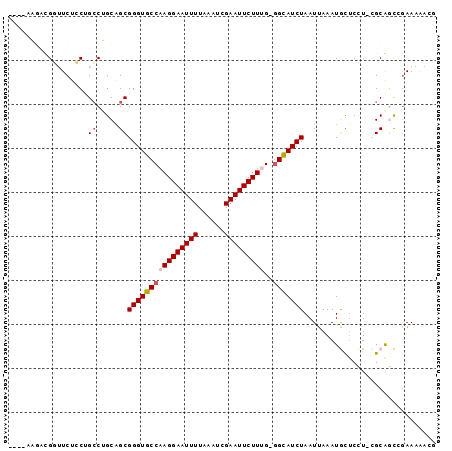
>2R_DroMel_CAF1 8411757 92 - 20766785 A---AAGACGGUUCUCCUGCCUGCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUG-GGCAUCUAAUUAAAUGCUCCU-CGCGGCAGAAAAACA .---......(((...(((((....(((((..(((((((((((......))))))))))-)((((........))))..))-))))))))...))). ( -29.30) >DroVir_CAF1 29362 89 - 1 ----GUCUUGCAUCUGC-GCCAGCGGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUCG-AGCAUCUAAUUAAAUACUCGU-AGCAUUCGAAUAUC- ----.((((((((((((-.......))))))).))))).........((((((.((.((-((.((........)).)))).-)).)))))).....- ( -25.50) >DroGri_CAF1 31607 92 - 1 --UGCUGUUGCAUCUGC-GCCAGCGGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUCGGGGCAUCUAAUUAAAUACGCGU-AGCAUUCGAAUAUC- --...((((((..((((-....)))).(((((((.((((((((......))))))))...)))))))............))-))))..........- ( -26.30) >DroEre_CAF1 25150 92 - 1 AA--AAGACGGUUCUCCUGCCUGCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUG-GGCAUCUAAUUAAAUGCUCCU-CGCGGCAGAAACA-G ..--.....((....))...((((.(((((..(((((((((((......))))))))))-)((((........))))..))-))).)))).....-. ( -29.20) >DroYak_CAF1 23207 92 - 1 A---AAGACGGUUCUCCUGCCUGCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUG-GGCAUCUAAUUAAAUGCUCCU-CGCGGCUGAAGAACG .---......(((((.(.(((....(((((..(((((((((((......))))))))))-)((((........))))..))-)))))).).))))). ( -30.80) >DroAna_CAF1 22842 87 - 1 ---------GUUUCUCCUGACUGCCGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUG-GGUAUCUAAUUAAAUGUUCCUUCGCAGCCAAAAGGUG ---------......(((....((.((((((((((((((((((......))))))))).-)))))))........(......))).))....))).. ( -21.30) >consensus ____AAGACGGUUCUCCUGCCUGCAGCGGGUGCCAAGGAAUUUUAAAUCGAAUUCUUUG_GGCAUCUAAUUAAAUGCUCCU_CGCAGCCGAAAAACG ..................((.....))((((((((((((((((......)))))))))..))))))).............................. (-14.51 = -15.03 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:46 2006