
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,408,226 – 8,408,318 |
| Length | 92 |
| Max. P | 0.968478 |

| Location | 8,408,226 – 8,408,318 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 77.06 |
| Mean single sequence MFE | -19.06 |
| Consensus MFE | -10.45 |
| Energy contribution | -11.05 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.601322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
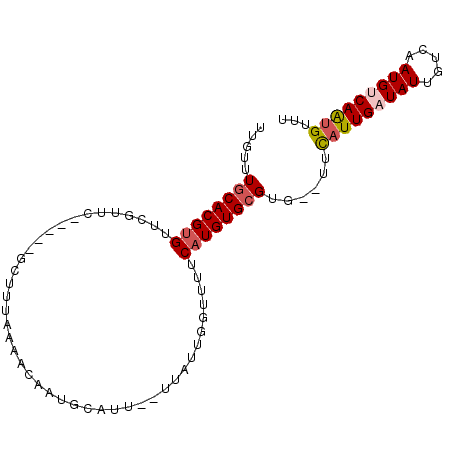
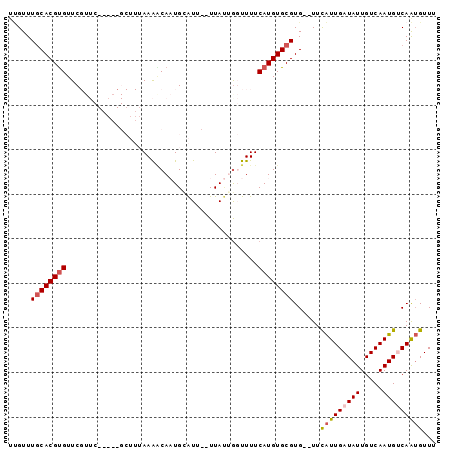
>2R_DroMel_CAF1 8408226 92 + 20766785 UUGUUUACACGCGUUCGUUCGAAACGCUUUAAAGCAAUGAAUU--UUAUUAGUUUUCAUGUGCGUG--UUCAUUGAUAUUGUCAAUGUCAAUGUUU ......((((((((((....).)))))......((((((((..--.........))))).))))))--).(((((((((.....)))))))))... ( -22.10) >DroSec_CAF1 19722 96 + 1 UUGUUUGCACGUGUUCGUUCGAAACGCUUUAAAGCAAUGCAUUUUUUAUGGGUUUUCAUGUGCGUGUGUUUAUUGAUAUUGUCAAUGUCAAUGUUU ......(((((((............(((....)))(((.(((.....))).)))..))))))).......(((((((((.....)))))))))... ( -20.20) >DroSim_CAF1 19352 91 + 1 GUGUUUGCACGUGUUCGUUC-----GCUUUAAAACAAUGCAUUUUUUAUGGGUUUUCAUGUGCGUGUUUUCAUUGAUAUUGUCAAUGUCAAUGUUU ......(((((((...(((.-----.......)))(((.(((.....))).)))..))))))).......(((((((((.....)))))))))... ( -19.20) >DroEre_CAF1 21653 76 + 1 U-GUUUGCACGUGUUC--------------AAGCCAUUGUGUU--UUA-UGCUUUUCAUGUGCGUG--UUCAUUGAUAUUGUCAAUGCCAGUGUUU .-....(((((((...--------------((((.((......--..)-)))))..)))))))...--..(((((((...)))))))......... ( -17.30) >DroYak_CAF1 19576 78 + 1 UCGUUUGCACGUGUUC--------------AAAACAUUGUGUU--UUACUGCUUUUCAUGUGCGUG--UUCGUUGAUAUUGUCAAUGACAGAGUUA ......(((((((...--------------(((((.....)))--)).........))))))).((--(.(((((((...))))))))))...... ( -16.50) >consensus UUGUUUGCACGUGUUCGUUC_____GCUUUAAAACAAUGCAUU__UUAUUGGUUUUCAUGUGCGUG__UUCAUUGAUAUUGUCAAUGUCAAUGUUU .....((((((((...........................................))))))))......(((((((((.....)))))))))... (-10.45 = -11.05 + 0.60)

| Location | 8,408,226 – 8,408,318 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 77.06 |
| Mean single sequence MFE | -14.28 |
| Consensus MFE | -6.11 |
| Energy contribution | -6.19 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.43 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968478 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
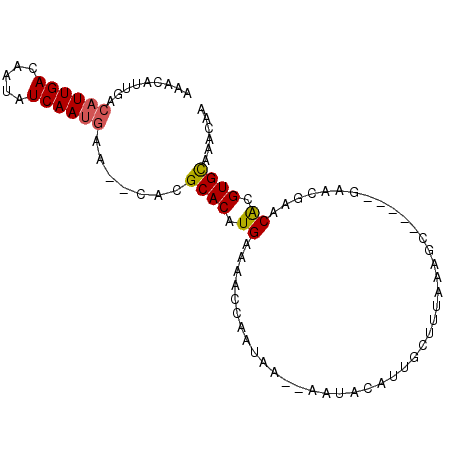
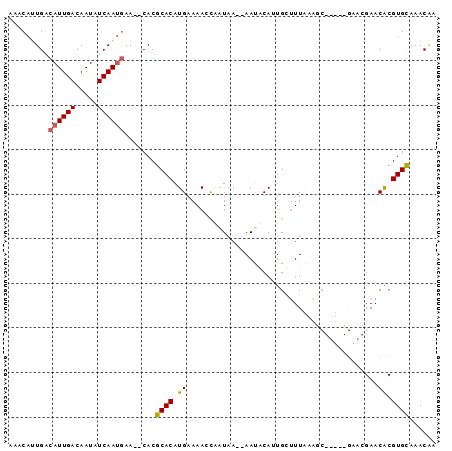
>2R_DroMel_CAF1 8408226 92 - 20766785 AAACAUUGACAUUGACAAUAUCAAUGAA--CACGCACAUGAAAACUAAUAA--AAUUCAUUGCUUUAAAGCGUUUCGAACGAACGCGUGUAAACAA .........((((((.....)))))).(--((((((.(((((.........--..))))))))......((((((.....))))))))))...... ( -20.00) >DroSec_CAF1 19722 96 - 1 AAACAUUGACAUUGACAAUAUCAAUAAACACACGCACAUGAAAACCCAUAAAAAAUGCAUUGCUUUAAAGCGUUUCGAACGAACACGUGCAAACAA ..........(((((.....)))))........(((((((......)))...((((((.((......))))))))...........))))...... ( -11.50) >DroSim_CAF1 19352 91 - 1 AAACAUUGACAUUGACAAUAUCAAUGAAAACACGCACAUGAAAACCCAUAAAAAAUGCAUUGUUUUAAAGC-----GAACGAACACGUGCAAACAC .........((((((.....)))))).......(((((((......)))..........((((((......-----))))))....))))...... ( -14.00) >DroEre_CAF1 21653 76 - 1 AAACACUGGCAUUGACAAUAUCAAUGAA--CACGCACAUGAAAAGCA-UAA--AACACAAUGGCUU--------------GAACACGUGCAAAC-A .........((((((.....))))))..--...((((.((..((((.-...--.........))))--------------...)).))))....-. ( -14.22) >DroYak_CAF1 19576 78 - 1 UAACUCUGUCAUUGACAAUAUCAACGAA--CACGCACAUGAAAAGCAGUAA--AACACAAUGUUUU--------------GAACACGUGCAAACGA ......((((.((((.....)))).).)--)).((((.((.((((((....--.......))))))--------------...)).))))...... ( -11.70) >consensus AAACAUUGACAUUGACAAUAUCAAUGAA__CACGCACAUGAAAACCAAUAA__AAUACAUUGCUUUAAAGC_____GAACGAACACGUGCAAACAA .........((((((.....)))))).......((((.((...........................................)).))))...... ( -6.11 = -6.19 + 0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:41 2006